AI developed in the UK is the world leader in identifying the location and expression of proteins
A new advanced artificial intelligence (AI) system has shown world-leading accuracy and speed in identifying protein patterns within individual cells. The new system, developed at the University of Surrey's Institute for People-Centred AI, could help scientists understand differences in cancer tumours and identify new drugs for diseases.
In a study published in Communications Biology, part of the Nature Portfolio, researchers demonstrate how the HCPL (Hybrid subCellular Protein Localiser) requires only partially labelled data to learn how to decipher the locations of proteins within cellular structures and their behaviour in different cells.
The team tested the HCPL on the Human Protein Atlas* and found it to be the most accurate tool for identifying the location of proteins within individual cells.
Professor Miroslaw Bober, leader of the HCPL project from the University of Surrey, said:
"To understand how proteins work inside cells, scientists need to study where they are located, but this can be a time-consuming and complicated process. HCPL makes this process easier.
"This program uses a deep-learning model to quickly and accurately identify subcellular structures where proteins are present inside individual cells. We are hopeful that HCPL can help scientists study how proteins work and develop new treatments for diseases."
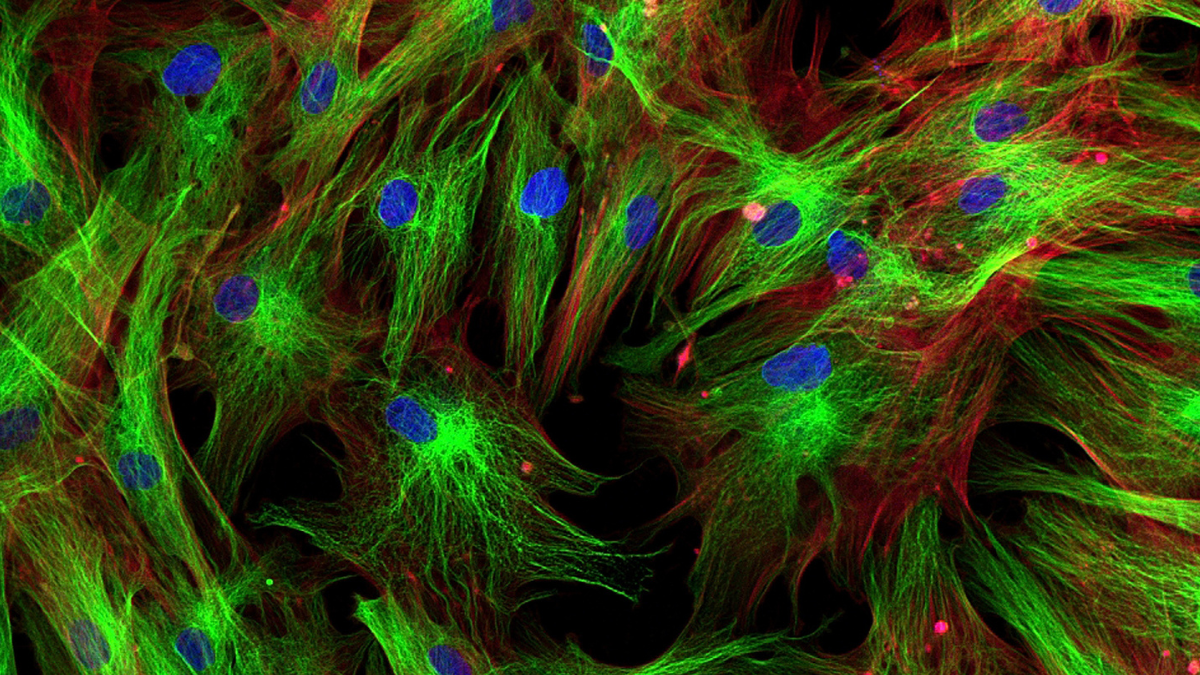
Spatial proteomics is a research area that studies the distribution of proteins in cells or tissues using a combination of experimental techniques and computational approaches. Fluorescence microscopy is a common method in this field where proteins are physically tagged with fluorescent markers. AI maps the proteins onto individual cell compartments (subcellular structures or organelles). This helps scientists to understand the roles and functions of proteins and possibly reveal the complex inner workings of cells.
HCPL was developed in partnership with ForecomAI, a research and development company with world-class expertise in machine and deep learning providing solutions in healthcare and biosciences.
Dr Amaia Irizar, director of ForecomAI said:
"Proteins play a key role in most cellular processes crucial to our survival. Unravelling protein distributions and interactions within individual cells is vital to understanding their functions and indispensable to developing new treatments.
"Our work with the University of Surrey enables scaling up of this process and opens new frontiers. The partnership between Surrey and ForecomAI has been a successful interdisciplinary collaboration in scientific research, paving the way for further initiatives."
The University of Surrey is a research-intensive university, producing world-leading research and delivering innovation in teaching to transform lives and change the world for the better. The University of Surrey's Institute for People-Centred AI combines over 30 years of technical excellence in the field of machine learning with multi-disciplinary research to answer the technical, ethical and governance questions that will enable the future of AI to be truly people-centred. A focus on research that makes a difference to the world has contributed to Surrey being ranked 55th in the world in the Times Higher Education (THE) University Impact Rankings 2022, which assesses more than 1,400 universities' performance against the United Nations' Sustainable Development Goals (SDGs).
###
- For more information, please contact the University of Surrey's press office via mediarelations@surrey.ac.uk.
- Professor Bober and his team are available for an interview upon request.
- *The Human Protein Atlas is a Swedish-based program initiated in 2003 with the aim to map all the human proteins in cells, tissues, and organs using an integration of various omics technologies, including antibody-based imaging, mass spectrometry-based proteomics, transcriptomics, and systems biology.
Paper details
- Husain, S.S., Ong, EJ., Minskiy, D. et al. Single-cell subcellular protein localisation using novel ensembles of diverse deep architectures. Commun Biol 6, 489 (2023). https://doi.org/10.1038/s42003-023-04840-z
Media Contacts
External Communications and PR team
Phone: +44 (0)1483 684380 / 688914 / 684378
Email: mediarelations@surrey.ac.uk
Out of hours: +44 (0)7773 479911
