Professor André Gerber
About
Biography
André studied Biochemistry at the University of Zurich and obtained a PhD in Cell Biology from the University of Basel for his work on adenosine deaminases acting on RNA. He completed post-doctoral studies in the laboratories of Dan Herschlag and Pat Brown at Stanford University, where he started to explore RNA-protein interactions on a genome-wide scale. During 2004-2011, he was an independent research group leader at the ETH Zurich, studying RNA-protein interactions with genomics and proteomics means in different model organisms. Since 2012 he is Professor of RNA biology at the Faculty of Health and Medical Sciences, University of Surrey, UK.
Research interests
We explore global and specific aspects of post-transcriptional gene regulation mediated by RNA-binding proteins and non-coding RNAs.
Gene expression must be tightly controlled to ensure coordinated synthesis of the cells’ macromolecular components. Besides transcriptional control, it has become evident that also the later post-transcriptional steps – namely the processing, transport, turnover and translation of mRNAs – play pivotal roles for diversification and spatiotemporal control of gene expression. Hundreds of RNA-binding proteins (RBPs) and non-coding RNAs mediate post-transcriptional control with significant implications in cell physiology and disease. Nevertheless, the targets and functions for most RNA-binding proteins and non-coding RNAs are not known.
We are combining genome-wide with classical biochemical and genetic tools to identify the RNA targets of RNA-binding proteins and to investigate post-transcriptional gene regulation on a global scale. Importantly, these studies revealed that RNA-binding proteins bind to and coordinate groups of mRNAs that code for proteins, which are localized to the same subcellular compartment, act in the same pathway or are components of macromolecular complexes, forming so-called RNA regulons. Moreover, these set of RNAs often bear conserved sequence/ structural elements that likely represent binding sites for RNA-binding proteins. These findings suggested the presence of a highly-organized post-transcriptional regulatory system that may affect virtually every RNA in a cell.
In our current research, we focus on specific RNA-binding proteins that coordinate the localization, decay or translation of mRNAs in the cytoplasm. Specfically, we investigate the functions of recently discovered 'unconventional' RNA-binding proteins, such as metabolic enzymes and aim to better understand the functional implications of those RNA-enzyme interactions. We also study how RBPs critically control expression of therapeutic mRNAs; and we develop models describing auto-regulatory feedback control through RNA-binding proteins. Furthermore, we characterize the 'translatome' – which refers to all mRNAs associated with ribosomes for protein synthesis (Halbeisen et al. 2009). We currently monitor the translatome and associated translational control during ageing and sleep in the brain and under pathological conditions. Commonly, we use budding yeast as model to establish new techniques and to elucidate principles of post-transcriptional control, and we work with mammalian cells to unravel the implications in disease.
For more information about our research and lab members, pls. consult our lab webpage (see link above; https://openwetware.org/wiki/Gerber).
Selected Reviews:
- Gerber, AP (2021) RNA-centric Approaches to Profile the RNA-protein Interaction Landscape on Selected RNAs. Non-coding RNA, 15(7(, 11.
- Albihlal, W.A., Gerber, A.P. (2018) Unconventional RNA-binding proteins: an uncharted zone in RNA biology. FEBS Lett. 592(17), 2917-2931.
- Iadevaia, V, Gerber, AP (2015) Combinatorial Control of mRNA Fates by RNA-Binding Proteins and Non-Coding RNAs. Biomolecules, 5(4):2207-22.
Publications
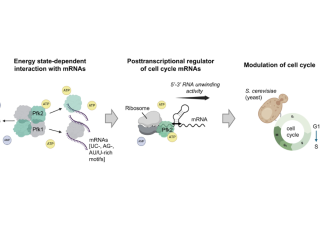
Phosphofructokinase (PFK) is a rate-limiting glycolytic enzyme that also possesses an unexplored RNA binding activity. Here, we show that the α-and β-subunits of yeast PFK, encoded by PFK1 and PFK2, respectively, bind hundreds of functionally related messenger RNAs (mRNAs) in cells, including one's coding for proteins involved in the regulation of mitotic cell cycle. Both Pfk1p and Pkf2p directly bind to short GA-, UC-, AU-, and U-rich motifs overrepresented in their mRNA targets. Strikingly, Pfk2p displays directional 5 –3 double-stranded RNA unwinding activity not seen with Pfk1p. Furthermore, Pfk2p dynamically associates with ribosomes and promotes translation of cell cycle genes. Consequently, pfk2Δ, but not pfk1Δ, mutants display increased cell sizes and severely delayed G1/S phase transition, independent of the enzyme's glycolytic activity. Our results uncovered a hidden function for the Pfk2 subunit as a translational activator of mitotic cell cycle gene transcripts possibly through energy-dependent RNA unwinding activity, thus providing a link between central energy metabolism and cell proliferation.
RNA-binding proteins are essential for gene regulation and the spatial organization of cells. Here, we report that the yeast ribosome biogenesis factor Loc1p is an intrinsically disordered RNA-binding protein with eight repeating positively charged, unstructured nucleic acid binding (PUN) motifs. While a single of these previously undefined motifs stabilizes folded RNAs, multiple copies strongly cooperate to catalyze RNA folding. In the presence of RNA, these multivalent PUN motifs drive phase separation. Proteome-wide searches in pro- and eukaryotes for proteins with similar arrays of PUN motifs reveal a strong enrichment in RNA-mediated processes and DNA remodeling. Thus, PUN motifs are potentially involved in a large variety of RNA- and DNA-related processes by concentrating them in membraneless organelles. The general function and wide distribution of PUN motifs across species suggest that in an ancient ‘RNA world’ PUN-like motifs may have supported the correct folding of early ribozymes.
ADAR1 and ADAR2 are members of a family of enzymes that catalyze the conversion of adenosine to inosine in double-stranded RNA. Unlike the other types of RNA editing that involve multiprotein editing complexes, the site-specific deamination of an adenosine to inosine is catalyzed by single enzymes. ADAR1 and ADAR2 have been purified and the genes cloned from various sources. Each gene encodes multiple splice variants. As it is crucial to have an adequate supply of pure protein to investigate this type of RNA editing, we describe in this article methods for both the purification and the overexpression of either full-length or partial ADAR1 and ADAR2 isoforms.
We describe a new, broadly applicable methodology for screening in parallel interactions of RNA-binding proteins (RBPs) with large numbers of microRNA (miRNA) precursors and for determining their affinities in native form in the presence of cellular factors. The assays aim at identifying pre-miRNAs that are potentially affected by the selected RBP during their biogenesis. The assays are carried out in microtiter plates and use chemiluminescent readouts. Detection of bound RBPs is achieved by protein or tag-specific antibodies allowing crude cell lysates to be used as a source of RBP. We selected 70 pre-miRNAs with phylogenetically conserved loop regions and 25 precursors of other well-characterized miRNAs for chemical synthesis in 3'-biotinylated form. An equivalent set in unmodified form served as inhibitors in affinity determinations. By testing three RBPs known to regulate miRNA biogenesis on this set of pre-miRNAs, we demonstrate that Lin28 and hnRNP A1 from cell lysates or as recombinant protein domains recognize preferentially precursors of the let-7 family, and that KSRP binds strongly to pre-miR-1-2.
The mammalian adenosine deaminases acting on RNA (ADARs) constitute a family of sequence-related proteins involved in pre-mRNA editing of nuclear transcripts through site-specific adenosine modification. We report here the identification and characterization of a human ADAR protein, hADAT1, that specifically deaminates adenosine 37 to inosine in eukaryotic tRNA Ala . It represents the functional homologue of the recently identified yeast protein Tad1p [Gerber, A., Grosjean, H., Melcher, T. & Keller, W. (1998) EMBO J. 17, 4780–4789]. The hADAT1 cDNA predicts a protein of 502 aa whose sequence displays strongest overall homology to a Drosophila melanogaster ORF (50% similarity, 32% identity), and the catalytic domain is closely related to the other ADAR proteins. In vitro , the recombinantly expressed and purified hADAT1 protein efficiently and specifically deaminates A 37 in the anticodon loop of tRNA Ala from higher eukaryotes and with lower efficiency from lower eukaryotes. It does not modify adenosines residing in double-stranded RNA or in pre-mRNAs that serve as substrates for ADAR1 or ADAR2. The anticodon stem–loop of tRNA Ala alone is not a functional substrate for hADAT1. The enzyme is expressed ubiquitously in human tissues and is represented by a single gene. The identification and cloning of hADAT1 should help to elucidate the physiological significance of this unique modification in tRNA Ala , which is conserved from yeast to man.
RNA-binding proteins (RBPs) have roles in the regulation of many post-transcriptional steps in gene expression, but relatively few RBPs have been systematically studied. We searched for the RNA targets of 40 proteins in the yeast Saccharomyces cerevisiae: a selective sample of the approximately 600 annotated and predicted RBPs, as well as several proteins not annotated as RBPs. At least 33 of these 40 proteins, including three of the four proteins that were not previously known or predicted to be RBPs, were reproducibly associated with specific sets of a few to several hundred RNAs. Remarkably, many of the RBPs we studied bound mRNAs whose protein products share identifiable functional or cytotopic features. We identified specific sequences or predicted structures significantly enriched in target mRNAs of 16 RBPs. These potential RNA-recognition elements were diverse in sequence, structure, and location: some were found predominantly in 3'-untranslated regions, others in 5'-untranslated regions, some in coding sequences, and many in two or more of these features. Although this study only examined a small fraction of the universe of yeast RBPs, 70% of the mRNA transcriptome had significant associations with at least one of these RBPs, and on average, each distinct yeast mRNA interacted with three of the RBPs, suggesting the potential for a rich, multidimensional network of regulation. These results strongly suggest that combinatorial binding of RBPs to specific recognition elements in mRNAs is a pervasive mechanism for multi-dimensional regulation of their post-transcriptional fate.
Post-transcriptional control of mRNAs by RNA-binding proteins (RBPs) has a prominent role in the regulation of gene expression. RBPs interact with mRNAs to control their biogenesis, splicing, transport, localization, translation, and stability. Defects in such regulation can lead to a wide range of human diseases from neurological disorders to cancer. Many RBPs are conserved between Caenorhabditis elegans and humans, and several are known to regulate apoptosis in the adult C. elegans germ line. How these RBPs control apoptosis is, however, largely unknown. Here, we identify mina-1(C41G7.3) in a RNA interference-based screen as a novel regulator of apoptosis, which is exclusively expressed in the adult germ line. The absence of MINA-1 causes a dramatic increase in germ cell apoptosis, a reduction in brood size, and an impaired P granules organization and structure. In vivo crosslinking immunoprecipitation experiments revealed that MINA-1 binds a set of mRNAs coding for RBPs associated with germ cell development. Additionally, a system-wide analysis of a mina-1 deletion mutant compared to wild type, including quantitative proteome and transcriptome data, hints to a post-transcriptional regulatory RBP network driven by MINA-1 during germ cell development in C. elegans. In particular, we found that the germline-specific Argonaute WAGO-4 protein levels are increased in mina-1 mutant background. Phenotypic analysis of double mutant mina-1;wago-4 revealed that contemporary loss of MINA-1 and WAGO-4 strongly rescues the phenotypes observed in mina-1 mutant background. To strengthen this functional interaction, we found that upregulation of WAGO-4 in mina-1 mutant animals causes hypersensitivity to exogenous RNAi. Our comprehensive experimental approach allowed us to describe a phenocritical interaction between two RBPs controlling germ cell apoptosis and exogenous RNAi. These findings broaden our understanding of how RBPs can orchestrate different cellular events such as differentiation and death in C. elegans.
RNA-binding proteins (RBPs) play pivotal roles in post-transcriptional regulation, acting as master controllers of the RNA’s life. The development of a variety of global analysis tools during the last decade has tremendously enriched our understanding of the function of RBPs and their implications in environmental and cellular cues. In this chapter, we discuss the properties and functions of the RBPs in fungi, and we outline the genome-wide approaches that have been developed during the last decade to systematically identify their RNA targets. We further sketch recent approaches aimed to search for novel RBPs, revealing that many more proteins such as metabolic enzymes could have additional functions in RNA regulation. Based on a rich dataset obtained in budding yeast, we further refer to attempts to integrate RNA targets for more than 60 RBPs into molecular networks to reveal structures and motifs. Finally, we elaborate on the future tasks and challenges on our way to complete our understanding of this highly connected and interwoven protein-RNA interaction network and its importance for cell physiology.
We report the characterization of tadA, the first prokaryotic RNA editing enzyme to be identified. Escherichia coli tadA displays sequence similarity to the yeast tRNA deaminase subunit Tad2p. Recombinant tadA protein forms homodimers and is sufficient for site-specific inosine formation at the wobble position (position 34) of tRNA(Arg2), the only tRNA having this modification in prokaryotes. With the exception of yeast tRNA(Arg), no other eukaryotic tRNA substrates were found to be modified by tadA. How ever, an artificial yeast tRNA(Asp), which carries the anticodon loop of yeast tRNA(Arg), is bound and modified by tadA. Moreover, a tRNA(Arg2) minisubstrate containing the anticodon stem and loop is sufficient for specific deamination by tadA. We show that nucleotides at positions 33-36 are sufficient for inosine formation in mutant Arg2 minisubstrates. The anticodon is thus a major determinant for tadA substrate specificity. Finally, we show that tadA is an essential gene in E.coli, underscoring the critical function of inosine at the wobble position in prokaryotes.
As our understanding of the complex network of regulatory pathways for gene expression continues to grow, avenues of investigation for how these new findings can be utilised in therapeutics are emerging. Recent growth of interest in the RNA binding protein (RBP) interactome has revealed it to be rich in targets linked to, and causative of diseases. While this is, in and of itself, very interesting, evidence is also beginning to arise for how the RBP interactome can act to modulate the response of diseases to existing therapeutic treatments, especially in cancers. Here we highlight this topic, providing examples of work that exemplifies such modulation of chemotherapeutic sensitivity.
Brain plasticity is induced by learning during wakefulness and is consolidated during sleep. But the molecular mechanisms involved are poorly understood and their relation to experience-dependent changes in brain activity remains to be clarified. Localised mRNA translation is important for the structural changes at synapses supporting brain plasticity consolidation. The translation mTOR pathway, via phosphorylation of 4E-BPs, is known to be activate during sleep and contributes to brain plasticity, but whether this activation is specific to synapses is not known. We investigated this question using acute exposure of rats to an enriched environment (EE). We measured brain activity with EEGs and 4E-BP phosphorylation at cortical and cerebellar synapses with Western blot analyses. Sleep significantly increased the conversion of 4E-BPs to their hyperphosphorylated forms at synapses, especially after EE exposure. EE exposure increased oscillations in the alpha band during active exploration and in the theta-to-beta (4–30 Hz) range, as well as spindle density, during NREM sleep. Theta activity during exploration and NREM spindle frequency predicted changes in 4E-BP hyperphosphorylation at synapses. Hence, our results suggest a functional link between EEG and molecular markers of plasticity across wakefulness and sleep.
RNA-binding proteins (RBPs) are key post-transcriptional regulators that play a substantial role during stress adaptation. Recent proteome-wide surveys have uncovered a large number of new and “unconventional” RBPs such as metabolic enzymes, yet little is known about the reconfiguration of the RNA-binding proteome (RBPome) and RNA-enzyme interactions in response to cellular stress. Here, we applied RNA-interactome capture to monitor the dynamics of the mRBPome upon mild oxidative stress in the yeast Saccharomyces cerevisiae. Among the 257 proteins that significantly changed RNA associations, we observed the coordinated remodeling of RNA-binding enzymes — particularly of the central carbon metabolism — that complemented known metabolic responses. Furthermore, we recognized the propensity for paralogous specific alterations of enzyme-RNA interactions. Our results suggest coordinated cross talk between RNA-enzyme interactions and intermediary metabolism to maintain the physiological and molecular balance upon oxidative stress, perhaps through specialization of paralogous during evolution.
Cardiac pathologies are characterized by intense remodeling of the extracellular matrix (ECM) that eventually leads to heart failure. Cardiomyocytes respond to the ensuing biomechanical stress by reexpressing fetal contractile proteins via transcriptional and posttranscriptional processes, such as alternative splicing (AS). Here, we demonstrate that the heterogeneous nuclear ribonucleoprotein C (hnRNPC) is up-regulated and relocates to the sarcomeric Z-disc upon ECM pathological remodeling. We show that this is an active site of localized translation, where the ribonucleoprotein associates with the translation machinery. Alterations in hnRNPC expression, phosphorylation, and localization can be mechanically determined and affect the AS of mRNAs involved in mechanotransduction and cardiovascular diseases, including Hippo pathway effector Yes-associated protein 1. We propose that cardiac ECM remodeling serves as a switch in RNA metabolism by affecting an associated regulatory protein of the spliceosome apparatus. These findings offer new insights on the mechanism of mRNA homeostatic mechanoregulation in pathological conditions.
Vascular endothelial growth factor A (VEGFA) plays a key role in the angiogenesis of human skin. Elevated levels of VEGFA are associated with several pathological conditions, including chronic inflammatory skin diseases and several types of skin cancer. In particular, squamous cell carcinoma (SCC) of the skin, the second most common skin cancer in the general population, is characterized by invasive growth, pronounced angiogenesis and elevated levels of VEGFA. The processing, turnover and production of VEGFA are extensively regulated at the post-transcriptional level, both by RNA-binding proteins and microRNAs (miRNAs). In the present study, we identified a new miRNA recognition element in a downstream conserved region of the VEGFA 3'-UTR. We confirmed the repressive effect of miR-361-5p on this element in vitro, identifying the first target for this miRNA. Importantly, we found that miR-361-5p levels are inversely correlated with VEGFA expression in SCC and in healthy skin, indicating that miR-361-5p could play a role in cancers.
Post-transcriptional control of gene expression is mediated by RNA-binding proteins (RBPs) and small non-coding RNAs (e.g., microRNAs) that bind to distinct elements in their mRNA targets. Here, we review recent examples describing the synergistic and/or antagonistic effects mediated by RBPs and miRNAs to determine the localisation, stability and translation of mRNAs in mammalian cells. From these studies, it is becoming increasingly apparent that dynamic rearrangements of RNA-protein complexes could have profound implications in human cancer, in synaptic plasticity, and in cellular differentiation.
On the 30th of March 2023, the RNA field lost one of its pioneers, Walter Keller, Professor Emeritus at the Biozentrum, University of Basel. Walter's exemplary and rigorous biochemistry work provided crucial tools for studying the entire life cycle of RNA. His dedication to the pursuit of robust evidence was inspiring for many, and helped drive forward several fields of research.
In the early years of Molecular Biology, the role of RNA was largely considered to be the transmission of the genetic information stored in the DNA into polypeptides. Three major types of RNA engaged in these processes were known: messenger RNA (mRNA), transfer RNA (tRNA) and ribosomal RNA (rRNA). However, within the last half century our knowledge about RNA classes, abundance and diversity of functions has increased dramatically. The discovery that the ribosome is essentially a ribozyme 1, 2 especially brought the central role of RNA for cellular life into the spotlight. The turn of the millennium was also the time of another far‐reaching realization, that of the pervasiveness of small RNA‐dependent regulation of gene expression 3. Within the decade that followed, RNAs were found in essentially all regulatory layers of gene expression, from the epigenetic layer at the top of the gene expression cascade 4, 5 to the most distal translation layer 6. Much of this rapid transition was enabled by sequencing technologies that were developed in the wake of the human genome project. Coupled with ingenious protocols for isolating RNAs of various sizes and molecular properties (e.g. 7) and computational methods to annotate various classes of small RNAs 8, the set of regulatory RNAs has expanded and diversified rapidly. We now know that < 2% of the human genome contains protein‐coding genes, whereas 80% is transcribed into noncoding RNAs (ncRNAs), classified based on their size into small and long noncoding RNAs (lncRNAs). Small ncRNAs include the highly investigated microRNAs (miRNAs) that were found to be increased in various cancer types where they exhibit pro‐oncogenic activity 9. LncRNAs are long (> 200 nucleotides) transcripts that functionally contribute to the control of cell differentiation and maintenance of cell identity. Notably, as most (~ 90%) of disease‐associated single nucleotide polymorphisms are located in gene regulatory or intergenic regions, long‐intergenic ncRNAs (lincRNAs) in particular may have pivotal impact for the development of personalized medicine therapies in the future10. Whether small or long, ncRNAs often bear domains enabling direct and specific interactions with other RNAs, forming RNA‐RNA hybrids that participate in the control of gene expression and biogenesis of RNAs 11.
MicroRNAs (miRNAs) are small, noncoding RNAs that negatively regulate gene expression. As miRNAs are involved in a wide range of biological processes and diseases, much effort has been invested in identifying their mRNA targets. Here, we present a novel combinatorial approach, RIP-chip-SRM (RNA-binding protein immunopurification + microarray + targeted protein quantification via selected reaction monitoring), to identify de novo high-confidence miRNA targets in the nematode Caenorhabditis elegans. We used differential RIP-chip analysis of miRNA-induced silencing complexes from wild-type and miRNA mutant animals, followed by quantitative targeted proteomics via selected reaction monitoring to identify and validate mRNA targets of the C. elegans bantam homolog miR-58. Comparison of total mRNA and protein abundance changes in mir-58 mutant and wild-type animals indicated that the direct bantam/miR-58 targets identified here are mainly regulated at the level of protein abundance, not mRNA stability.
Genes encoding RNA-binding proteins are diverse and abundant in eukaryotic genomes. Although some have been shown to have roles in post-transcriptional regulation of the expression of specific genes, few of these proteins have been studied systematically. We have used an affinity tag to isolate each of the five members of the Puf family of RNA-binding proteins in Saccharomyces cerevisiae and DNA microarrays to comprehensively identify the associated mRNAs. Distinct groups of 40-220 different mRNAs with striking common themes in the functions and subcellular localization of the proteins they encode are associated with each of the five Puf proteins: Puf3p binds nearly exclusively to cytoplasmic mRNAs that encode mitochondrial proteins; Puf1p and Puf2p interact preferentially with mRNAs encoding membrane-associated proteins; Puf4p preferentially binds mRNAs encoding nucleolar ribosomal RNA-processing factors; and Puf5p is associated with mRNAs encoding chromatin modifiers and components of the spindle pole body. We identified distinct sequence motifs in the 3'-untranslated regions of the mRNAs bound by Puf3p, Puf4p, and Puf5p. Three-hybrid assays confirmed the role of these motifs in specific RNA-protein interactions in vivo. The results suggest that combinatorial tagging of transcripts by specific RNA-binding proteins may be a general mechanism for coordinated control of the localization, translation, and decay of mRNAs and thus an integral part of the global gene expression program.
Several transfer RNAs (tRNAs) contain inosine (I) at the first position of their anticodon (position 34); this modification is thought to enlarge the codon recognition capacity during protein synthesis. The tRNA-specific adenosine deaminase of Saccharomyces cerevisiae that forms I(34) in tRNAs is described. The heterodimeric enzyme consists of two sequence-related subunits (Tad2p/ADAT2 and Tad3p/ADAT3), both of which contain cytidine deaminase (CDA) motifs. Each subunit is encoded by an essential gene (TAD2 and TAD3), indicating that I(34) is an indispensable base modification in elongating tRNAs. These results provide an evolutionary link between the CDA superfamily and RNA-dependent adenosine deaminases (ADARs/ADATs).
RNA-bindingproteins (RBPs) play important roles in the posttranscriptional control of gene expression. However, our understanding of how RBPs interact with each other at different regulatory levels to coordinate the RNA metabolism of the cell is rather limited. Here, we construct the posttranscriptional regulatory network among 69 experimentally studied RBPs in yeast to show that more than one-third of the RBPs autoregulate their expression at the posttranscriptional level and demonstrate that autoregulatory RBPs show reduced protein noise with a tendency to encode for hubs in this network. We note that in- and outdegrees in the posttranscriptional RBP–RBP regulatory network exhibit gaussian and scale-free distributions, respectively. This network was also densely interconnected with extensive cross-talk between RBPs belonging to different posttranscriptional steps, regulating varying numbers of cellular RNA targets. We show that feed-forward loops and superposed feed-forward/feedback loops are the most significant three-node subgraphs in this network. Analysis of the corresponding protein–proteininteraction (posttranslational) network revealed that it is more modular than the posttranscriptional regulatory network. There is significant overlap between the regulatory and protein–proteininteraction networks, with RBPs that potentially control each other at the posttranscriptional level tending to physically interact and being part of the same ribonucleoprotein (RNP) complex. Our observations put forward a model wherein RBPs could be classified into those that can stably interact with a limited number of protein partners, forming stable RNP complexes, and others that form transient hubs, having the ability to interact with multiple RBPs forming many RNPs in the cell.
Pre-mRNA editing involving the conversion of adenosine to inosine is mediated by adenosine deaminases that act on RNA (ADAR1 and ADAR2). ADARs contain multiple double-stranded RNA(dsRNA)-binding domains in addition to an adenosine deaminase domain. An adenosine deaminase acting on tRNAs, scTad1p (also known as scADAT1), cloned from Saccharomyces cerevisiae has a deaminase domain related to the ADARs but lacks dsRNA-binding domains. We have identified a gene homologous to scADAT1 in the region of Drosophila melanogaster Adh chromosome II. Recombinant Drosophila ADAT1 (dADAT1) has been expressed in the yeast Pichia pastoris and purified. The enzyme has no activity on dsRNA substrates but is a tRNA deaminase with specificity for adenosine 37 of insect alanine tRNA. dADAT1 shows greater similarity to vertebrate ADARs than to yeast Tad1p, supporting the hypothesis of a common evolutionary origin for ADARs and ADATs. dAdat1 transcripts are maternally supplied in the egg. Zygotic expression is widespread initially and later concentrates in the central nervous system.
In vivo characterization of RNA-protein interactions is the key for understanding RNA regulatory mechanisms. Herein, we describe a protocol for detection of proteins interacting with polyadenylated RNAs in the yeast Saccharomyces cerevisiae. Proteins are crosslinked to nucleic acids in vivo by ultraviolet (UV) irradiation of cells, and poly(A)-containing RNAs with bound proteins are isolated from cell lysates using oligo[dT]25 beads. RBPs can be detected by immunoblot analysis or with mass spectrometry to define the mRNA-binding proteome (mRBPome) and its changes under stress.
Some of the classical paradigms of gene regulation have been challenged by global-scale analysis of eukaryotic transcriptional and post-transcriptional gene regulation (PTGR), made possible by the development of genomics and proteomics tools. Post-transcriptional events in particular are increasingly being recognized as important sources of gene regulation. The hundreds of regulatory RNA-binding proteins that exist in eukaryotes may regulate dozens to hundreds of functionally related RNA targets. Likewise, the expression of considerable fractions of many eukaryotic genomes is affected by hundreds of non-coding RNAs, e.g., microRNAs. These findings suggest an enormous regulatory potential for PTGR that may affect virtually every message in a cell. All gene regulatory systems are composed of simple network circuits that coordinate the transfer of regulatory signals to a target gene/message.
Homeostatic regulation of ionic currents is of paramount importance during periods of synaptic growth or remodeling. Our previous work has identified the translational repressor Pumilio (Pum) as a regulator of sodium current (I(Na)) and excitability in Drosophila motoneurons. In this current study, we show that Pum is able to bind directly the mRNA encoding the Drosophila voltage-gated sodium channel paralytic (para). We identify a putative binding site for Pum in the 3' end of the para open reading frame (ORF). Characterization of the mechanism of action of Pum, using whole-cell patch clamp and real-time reverse transcription-PCR, reveals that the full-length protein is required for translational repression of para mRNA. Additionally, the cofactor Nanos is essential for Pum-dependent para repression, whereas the requirement for Brain Tumor (Brat) is cell type specific. Thus, Pum-dependent regulation of I(Na) in motoneurons requires both Nanos and Brat, whereas regulation in other neuronal types seemingly requires only Nanos but not Brat. We also show that Pum is able to reduce the level of nanos mRNA and as such identify a potential negative-feedback mechanism to protect neurons from overactivity of Pum. Finally, we show coupling between I(Na) (para) and I(K) (Shal) such that Pum-mediated change in para results in a compensatory change in Shal. The identification of para as a direct target of Pum represents the first ion channel to be translationally regulated by this repressor and the location of the binding motif is the first example in an ORF rather than in the canonical 3'-untranslated region of target transcripts.
Whilst the profiling of the transcriptome and proteome even of single-cells becomes feasible, the analysis of the translatome, which refers to all messenger RNAs (mRNAs) engaged with ribosomes for protein synthesis, is still an elaborate procedure requiring millions of cells. Herein, we report the generation and use of “smart materials”, namely molecularly imprinted polymers (MIPs) to facilitate the isolation of ribosomes and translated mRNAs from merely 1,000 cells. In particular, we show that a hydrogel-based ribosome imprinted polymer could recover ribosomes and associated mRNAs from human, simian and mice cellular extracts, but did not selectively enrich yeast ribosomes, thereby demonstrating selectivity. Furthermore, ribosome imprinted polymers enabled the sensitive measurement of an mRNA translational regulatory event, requiring 1,000-fold less cells than current methodologies. These results provide first evidence for the suitability of MIPs to selectively recover ribonucleoprotein complexes such as ribosomes, founding a novel means for sensitive detection of gene regulation.
RNA–protein interactions frame post-transcriptional regulatory networks and modulate transcription and epigenetics. While the technological advances in RNA sequencing have significantly expanded the repertoire of RNAs, recently developed biochemical approaches combined with sensitive mass-spectrometry have revealed hundreds of previously unrecognized and potentially novel RNA-binding proteins. Nevertheless, a major challenge remains to understand how the thousands of RNA molecules and their interacting proteins assemble and control the fate of each individual RNA in a cell. Here, I review recent methodological advances to approach this problem through systematic identification of proteins that interact with particular RNAs in living cells. Thereby, a specific focus is given to in vivo approaches that involve crosslinking of RNA–protein interactions through ultraviolet irradiation or treatment of cells with chemicals, followed by capture of the RNA under study with antisense-oligonucleotides and identification of bound proteins with mass-spectrometry. Several recent studies defining interactomes of long non-coding RNAs, viral RNAs, as well as mRNAs are highlighted, and short reference is given to recent in-cell protein labeling techniques. These recent experimental improvements could open the door for broader applications and to study the remodeling of RNA–protein complexes upon different environmental cues and in disease.
The regulation of mRNA translation plays an essential role in neurons, contributing to important brain functions, such as brain plasticity and memory formation. Translation is conducted by ribosomes, which at their core consist of ribosomal proteins (RPs) and ribosomal RNAs. While translation can be regulated at diverse levels through global or mRNA-specific means, recent evidence suggests that ribosomes with distinct configurations are involved in the translation of different subsets of mRNAs. However, whether and how such proclaimed ribosome heterogeneity could be connected to neuronal functions remains largely unresolved. Here, we postulate that the existence of heterologous ribosomes within neurons, especially at discrete synapses, subserve brain plasticity. This hypothesis is supported by recent studies in rodents showing that heterogeneous RP expression occurs in dendrites, the compartment of neurons where synapses are made. We further propose that sleep, which is fundamental for brain plasticity and memory formation, has a particular role in the formation of heterologous ribosomes, specialised in the translation of mRNAs specific for synaptic plasticity. This aspect of our hypothesis is supported by recent studies showing increased translation and changes in RP expression during sleep after learning. Thus, certain RPs are regulated by sleep, and could support different sleep functions, in particular brain plasticity. Future experiments investigating cell-specific heterogeneity in RPs across the sleep-wake cycle and in response to different behaviour would help address this question
RNA-binding proteins (RBPs) play key roles in the post-transcriptional control of gene expression. Therefore, biochemical characterization of mRNA-protein complexes is essential to understanding mRNA regulation inferred by interacting proteins or non-coding RNAs. Herein, we describe a tandem RNA isolation procedure (TRIP) that enables the purification of endogenously formed mRNA-protein complexes from cellular extracts. The two-step protocol involves the isolation of polyadenylated mRNAs with antisense oligo(dT) beads and subsequent capture of an mRNA of interest with 3'-biotinylated 2'-O-methylated antisense RNA oligonucleotides, which can then be isolated with streptavidin beads. TRIP was used to recover in vivo crosslinked mRNA-ribonucleoprotein (mRNP) complexes from yeast, nematodes and human cells for further RNA and protein analysis. Thus, TRIP is a versatile approach that can be adapted to all types of polyadenylated RNAs across organisms to study the dynamic re-arrangement of mRNPs imposed by intracellular or environmental cues.
Trf4p and Trf5p are non-canonical poly(A polymerases and are part of the heteromeric protein complexes TRAMP4 and TRAMP5 that promote the degradation of aberrant and short-lived RNA substrates by interacting with the nuclear exosome. To assess the level of functional redundancy between the paralogous Trf4 and Trf5 proteins and to investigate the role of the Trf4-dependent polyadenylation in vivo, we used DNA microarrays to compare gene expression of the wild-type yeast strain of S. cerevisiae with either that of trf4D or trf5D mutant strains or the trf4D mutant expressing the polyadenylationdefective Trf4(DADA) protein. We found little overlap between the sets of transcripts with altered expression in the trf4D or the trf5D mutants, suggesting that Trf4p and Trf5p target distinct groups of RNAs for degradation. Surprisingly, most RNAs the expression of which was altered by the trf4 deletion were restored to wild-type levels by overexpression of TRF4(DADA), showing that the polyadenylation activity of Trf4p is dispensable in vivo. Apart from previously reported Trf4p and Trf5p target RNAs, this analysis along with in vivo cross-linking and RNA immunopurification-chip experiments revealed that both the TRAMP4 and the TRAMP5 complexes stimulate the degradation of spliced-out introns via a mechanism that is independent of the polyadenylation activity of Trf4p. In addition, we show that disruption of trf4 causes severe shortening of telomeres suggesting that TRF4 functions in the maintenance of telomere length. Finally, our study demonstrates that TRF4, the exosome, and TRF5 participate in antisense RNA–mediated regulation of genes involved in phosphate metabolism. In conclusion, our results suggest that paralogous TRAMP complexes have distinct RNA selectivities with functional implications in RNA surveillance as well as other RNA–related processes. This indicates widespread and integrative functions of TRAMP complexes for the coordination of different gene expression regulatory processes.
The double-stranded RNA-specific adenosine deaminases ADAR1 and ADAR2 convert adenosine (A) residues to inosine (I) in messenger RNA precursors (pre-mRNA). Their main physiological substrates are pre-mRNAs encoding subunits of ionotropic glutamate receptors or serotonin receptors in the brain. ADAR1 and ADAR2 have similar sequence features, including double-stranded RNA binding domains (dsRBDs) and a deaminase domain. The tRNA-specific adenosine deaminases Tad1p and Tad2p/Tad3p modify A 37 in tRNA-Ala1 of eukaryotes and the first nucleotide of the anticodon (A 34) of several bacterial and eukaryotic tRNAs, respectively. Tad1p is related to ADAR1 and ADAR2 throughout its sequence but lacks dsRBDs. Tad1p could be the ancestor of ADAR1 and ADAR2. The deaminase domains of ADAR1, ADAR2 and Tad1p are very similar and resemble the active site domains of cytosine/cytidine deaminases.
Post-transcriptional regulation of gene expression plays important roles in diverse cellular processes such as development, metabolism and cancer progression. Whereas many classical studies explored the mechanistics and physiological impact on specific mRNA substrates, the recent development of genome-wide analysis tools enables the study of post-transcriptional gene regulation on a global scale. Importantly, these studies revealed distinct programs of RNA regulation, suggesting a complex and versatile post-transcriptional regulatory network. This network is controlled by specific RNA-binding proteins and/or non-coding RNAs, which bind to specific sequence or structural elements in the RNAs and thereby regulate subsets of mRNAs that partly encode functionally related proteins. It will be a future challenge to link the spectra of targets for RNA-binding proteins to post-transcriptional regulatory programs and to reveal its physiological implications.
We describe ribosome affinity purification (RAP), a method that allows rapid purification of ribosomes and associated messages from the yeast Saccharomyces cerevisiae. The method relies on the expression of protein A tagged versions of the ribosomal protein Rpl16, which is used to efficiently recover endogenously formed ribosomes and polysomes from cellular extracts with IgG-coupled spherical microbeads. This approach can be applied to profile reactions of the translatome, which refers to all messages associated with ribosomes, with those of the transcriptome using DNA microarrays. In addition, ribosomal proteins, their modifications, and/or other associated proteins can be mapped with mass spectrometry. Finally, application of this method in other organisms provides a valuable tool to decipher cell-type specific gene expression patterns.
Post-transcriptional gene regulation is largely mediated by RNA-binding proteins (RBPs) that modulate mRNA expression at multiple levels, from RNA processing to translation, localization, and degradation. Thereby, the genome-wide identification of mRNAs regulated by RBPs is crucial to uncover post--transcriptional gene regulatory networks. In this chapter, we provide a detailed protocol for one of the techniques that has been developed to systematically examine RNA targets for RBPs. This technique involves the purification of endogenously formed RBP-mRNA complexes with specific antibodies from cellular extracts, followed by the identification of associated RNAs using DNA microarrays. Such RNA-binding protein immunopurification-microarray profiling, also called RIP-Chip, has also been applied to identify mRNAs that are transported to distinct subcellular compartments by RNP-motor complexes. The application and further development of this method could provide global insights into the subcellular architecture of the RBP-RNA network, and how it is restructured upon changing environmental conditions, during development, and possibly in disease.
RNA-binding proteins (RBPs) participate in several steps of post-transcriptional regulation of gene expression, such as splicing, messenger RNA transport, mRNA localization, and translation. Gene-expression regulation in trypanosomatids occurs primarily at the post-transcriptional level, and RBPs play important roles in the process. Here, we characterized the RBP TcSgn1, which contains one RNA recognition motif (RRM). TcSgn1 is a close ortholog of yeast Saccharomyces cerevisiae protein ScSgn1, which plays a role in translational regulation in the cytoplasm. We found that TcSgn1 in Trypanosoma cruzi is localized in the nucleus in exponentially growing epimastigotes. By performing immunoprecipitation assays of TcSgn1, we identified hundreds of mRNAs associated with the protein, a significant fraction of them coding for nucleic acids binding, transcription, and endocytosis proteins. In addition, we show that TcSgn1 is capable of interacting directly with the poly(A) tail of the mRNAs. The study of parasites under nutritional stress showed that TcSgn1 was localized in cytoplasmic granules in addition to localizing in the nucleus. Similar to ScSgn1, we observed that TcSgn1 also interacts with the PABP1 protein, suggesting that this protein may play a role in regulating gene expression in T. cruzi. Taken together, our results show that RNA-binding protein TcSgn1 is part of ribonucleoprotein complexes associated with nuclear functions, stress response, and RNA metabolism.
Cells rapidly alter gene expression in response to environmental stimuli such as nutrients, hormones, and drugs. During the imposed ‘‘remodeling’’ of gene expression, changes in the levels of particular mRNAs do not necessarily correlate with those of the encoded proteins, which could in part rely on the differential recruitment of mRNAs to translating ribosomes. To systematically address this issue, we have established an approach to rapidly access the translational status of each mRNA in the yeast Saccharomyces cerevisiae by affinity purification of endogenously formed ribosomes and the analysis of associated mRNAs with DNA microarrays. Using this method, we compared changes in total mRNA levels (transcriptome) with ribosome associations (translatome) after the application of different conditions of cellular stress. Severe stresses, induced by amino acid depletion or osmotic shock, stimulated highly correlated responses affecting about 15% of both total RNA levels and translatome. Many of the regulated messages code for functionally related proteins, thus reflecting logical responses to the particular stress. In contrast, mild stress provoked by addition of Calcofluor-white and menadione altered the translatome of approximately 1% of messages with only marginal effects on total mRNA, suggesting largely uncorrelated responses of transcriptome and translatome. Among these putative translationally regulated messages were most components of the mitochondrial ATPase. Increased polysome associations of corresponding messages and higher mitochondrial ATPase activities upon treatment confirmed the relevance for regulation of this macromolecular complex. Our results suggest the presence of highly sensitive translational regulatory networks that coordinate functionally related messages. These networks are preferentially activated for rapid adaptation of cells to minor environmental perturbations.
Cytoplasmic mRNA localization provides a means of generating cell asymmetry and segregating protein activity. Previous studies have identified two mRNAs that localize to the bud tips of the yeast Saccharomyces cerevisiae. To identify additional localized mRNAs, we immunoprecipitated the RNA transport components She2p, She3p, and Myo4p and performed DNA microarray analysis of their associated RNAs. A secondary screen, using a GFP-tagged RNA reporter assay, identified 22 mRNAs that are localized to bud tips. These messages encode a wide variety of proteins, including several involved in stress responses and cell wall maintenance. Many of these proteins are asymmetrically localized to buds. However, asymmetric localization also occurs in the absence of RNA transport, suggesting the existence of redundant protein localization mechanisms. In contrast to findings in metazoans, the untranslated regions are dispensable for mRNA localization in yeast. This study reveals an unanticipated widespread use of RNA transport in budding yeast.
The La-motif (LAM) is an ancient and ubiquitous RNA-binding domain defining a superfamily of proteins, which comprises the genuine La proteins and La-related proteins (LARPs). In contrast to La, which binds and stabilizes pre-tRNAs and other RNA polymerase III transcripts, data on function and RNA targets of the LARPs have remained scarce. We have undertaken a global approach to elucidate the previously suggested role of the yeast LARP Slf1p in copper homeostasis. By applying RNA-binding protein immunopurification-microarray (RIP-Chip) analysis, we show that Slf1p and its paralog Sro9p copurify with overlapping sets of hundreds of functionally related mRNAs, including many transcripts coding for ribosomal proteins and histones. Interestingly, among these potential RNA targets were also mRNAs coding for proteins critical for protection of cells against elevated copper concentrations. Mutations introduced in the conserved aromatic patch of the LAM in Slf1p drastically impaired both association with its targets and Slf1-mediated protection of cells against toxic copper concentrations. Furthermore, we show that Slf1p stabilizes copper-related mRNA targets in a LAM-dependent manner. These results provide the first evidence for post-transcriptional regulation of factors/pathways implicated in copper homeostasis by a cytoplasmic RBP.
Gene expression is regulated at multiple levels to ensure coordinated synthesis of the cells’ macromolecular components. Besides transcriptional regulation, the control of the later post-transcriptional steps has substantial impact on gene expression with widespread implications in physiologically important processes such as development, metabolism, neuronal function, and for cancer progression. On the one hand, posttranscriptional regulation is mediated by RNA-binding proteins (RBPs), which control almost every aspect of RNA’s life in a dynamic manner from RNA maturation, quality control, localization, translation, and degradation. On the other hand, mRNAs are post-transcriptionally regulated via physical interactions with small non-coding RNAs. The best characterized class of such RNAs are microRNAs (miRNAs), ~22 nucleotide long RNA molecules that negatively regulate gene expression. Hundreds of RNA-binding proteins (RBPs) and miRNAs are present in eukaryotic genomes, rivaling in number other classes of regulatory molecules such as transcription factors and kinases and thus, suggests and elaborate system for post-transcriptional control that may affect virtually every message in a cell. Whereas many classical studies explored the molecular function and physiological impact of post-transcriptional regulation on specific mRNA substrates, the recent development of genome-wide analysis tools enables now to study the extend and logic of post-transcriptional gene regulation (PTGR) on a global scale. I have pioneered such ‘ribonomic’ studies and established methods to affinity isolate RNA-binding proteins and systematically analyzed bound RNAs with DNA microarrays for more than 50 RBPs from yeast, flies, and humans. These studies revealed that RBPs preferentially associate with messages that share common functional and structural attributes suggesting the presence of a highly complex and interweaved post-transcriptional regulatory system. In addition, unraveling the RNA targets for particular RBPs has lead to new insights into their molecular and physiological function. In this habilitation thesis, I summarize some of these investigations and provide an outlook for future research and potential applications for pharmaceutical sciences.
Genome-wide identification of mRNAs regulated by RNA-binding proteins is crucial to uncover post-transcriptional gene regulatory systems. The conserved PUF family RNA-binding proteins repress gene expression post-transcriptionally by binding to sequence elements in 3'-UTRs of mRNAs. Despite their well-studied implications for development and neurogenesis in metazoa, the mammalian PUF family members are only poorly characterized and mRNA targets are largely unknown. We have systematically identified the mRNAs associated with the two human PUF proteins, PUM1 and PUM2, by the recovery of endogenously formed ribonucleoprotein complexes and the analysis of associated RNAs with DNA microarrays. A largely overlapping set comprised of hundreds of mRNAs were reproducibly associated with the paralogous PUM proteins, many of them encoding functionally related proteins. A characteristic PUF-binding motif was highly enriched among PUM bound messages and validated with RNA pull-down experiments. Moreover, PUF motifs as well as surrounding sequences exhibit higher conservation in PUM bound messages as opposed to transcripts that were not found to be associated, suggesting that PUM function may be modulated by other factors that bind conserved elements. Strikingly, we found that PUF motifs are enriched around predicted miRNA binding sites and that high-confidence miRNA binding sites are significantly enriched in the 3'-UTRs of experimentally determined PUM1 and PUM2 targets, strongly suggesting an interaction of human PUM proteins with the miRNA regulatory system. Our work suggests extensive connections between the RBP and miRNA post-transcriptional regulatory systems and provides a framework for deciphering the molecular mechanism by which PUF proteins regulate their target mRNAs.
Aging is associated with substantial physiological changes and constitutes a major risk factor for neurological disorders including dementia. Alterations in gene expression upon aging have been extensively studied; however, an in-depth characterization of post-transcriptional regulatory events remains elusive. Here, we profiled the age-related changes of the transcriptome and translatome in the female mouse hippocampus by RNA sequencing of total RNA and polysome preparations at four ages (3-, 6-, 12-, 20-month-old); and we implemented a variety of bioinformatics approaches to unravel alterations in transcript abundance, alternative splicing, and polyadenylation site selection. We observed mostly well-coordinated transcriptome and translatome expression signatures across age including upregulation of transcripts related to immune system processes and neuroinflammation, though transcripts encoding ribonucleoproteins or associated with mitochondrial functions, calcium signaling and the cell-cycle displayed substantial discordant profiles, suggesting translational control associated with age-related deficits in hippocampal-dependent behavior. By contrast, alternative splicing was less preserved, increased with age and was associated with distinct functionally-related transcripts encoding proteins acting at synapses/dendrites, RNA-binding proteins; thereby predicting regulatory roles for RBM3 and CIRBP. Only minor changes in polyadenylation site selection were identified, indicating pivotal 3′-end selection in young adults compared to older groups. Overall, our study provides a comprehensive resource of age-associated post-transcriptional regulatory events in the mouse hippocampus, enabling further examination of the molecular features underlying age-associated neurological diseases.
During the past decade, there has been a rapidly increased appreciation of the role of translation as a key regulatory node in gene expression. Thereby, the development of methods to infer the translatome, which refers to the entirety of mRNAs associated with ribosomes for protein synthesis, has facilitated the discovery of new principles and mechanisms of translation and expanded our view of the underlying logic of protein synthesis. Here, we review the three main methodologies for translatome analysis, and we highlight some of the recent discoveries made using each technique. We first discuss polysomal profiling, a classical technique that involves the separation of mRNAs depending on the number of bound ribosomes using a sucrose gradient, and which has been combined with global analysis tools such as DNA microarrays or high-throughput RNA sequencing to identify the RNAs in polysomal fractions. We then introduce ribosomal profiling, a recently established technique that enables the mapping of ribosomes along mRNAs at near-nucleotide resolution on a global scale. We finally refer to ribosome affinity purification techniques that are based on the cell-type-specific expression of tagged ribosomal proteins, allowing the capture of translatomes from specialized cells in organisms. We discuss the advantages and disadvantages of these three main techniques in the pursuit of defining the translatome, and we speculate about future developments.
mRNAs encoding mitochondrial proteins are enriched in the vicinity of mitochondria, presumably to facilitate protein transport. A possible mechanism for enrichment may involve interaction of the translocase of the mitochondrial outer membrane (TOM) complex with the precursor protein while it is translated, thereby leading to association of polysomal mRNAs with mitochondria. To test this hypothesis, we isolated mitochondrial fractions from yeast cells lacking the major import receptor, Tom20, and compared their mRNA repertoire to that of wild-type cells by DNA microarrays. Most mRNAs encoding mitochondrial proteins were less associated with mitochondria, yet the extent of decrease varied among genes. Analysis of several mRNAs revealed that optimal association of Tom20 target mRNAs requires both translating ribosomes and features within the encoded mitochondrial targeting signal. Recently, Puf3p was implicated in the association of mRNAs with mitochondria through interaction with untranslated regions. We therefore constructed a tom20Δ puf3Δ double-knockout strain, which demonstrated growth defects under conditions where fully functional mitochondria are required. Mislocalization effects for few tested mRNAs appeared stronger in the double knockout than in the tom20Δ strain. Taken together, our data reveal a large-scale mRNA association mode that involves interaction of Tom20p with the translated mitochondrial targeting sequence and may be assisted by Puf3p.
The posttranscriptional modification of messenger RNA precursors (pre-mRNAs) by base deamination can profoundly alter the physiological function of the encoded proteins. The recent identification of tRNA-specific adenosine deaminases (ADATs) has led to the suggestion that these enzymes, as well as the cytidine and adenosine deaminases acting on pre-mRNAs (CDARs and ADARs), belong to a superfamily of RNA-dependent deaminases. This superfamily might have evolved from an ancient cytidine deaminase. This article reviews the reactions catalysed by these enzymes and discusses their evolutionary relationships.
Post-transcriptional control of gene expression is mediated via RNA-binding proteins (RBPs) that interact with mRNAs in a combinatorial fashion. While recent global RNA interactome capture experiments expanded the repertoire of cellular RBPs quiet dramatically, little is known about the assembly of RBPs on particular mRNAs; and how these associations change and control the fate of the mRNA in drug-treatment conditions. Here we introduce a novel biochemical approach, termed tobramycinbased tandem RNA isolation procedure (tobTRIP), to quantify proteins associated with the 3'UTRs of cyclin-dependent kinase inhibitor 1B (CDKN1B/p27Kip1) mRNAs in vivo. P27Kip1 plays an important role in mediating a cell's response to cisplatin (CP), a widely used chemotherapeutic cancer drug that induces DNA damage and cell cycle arrest. We found that p27Kip1 mRNA is stabilized upon CP treatment of HEK293 cells through elements in its 3'UTR. Applying tobTRIP, we further compared the associated proteins in CP and non-treated cells, and identified more than fifty interacting RBPs, many functionally related and evoking a coordinated response. Knock-downs of several of the identified RBPs in HEK293 cells confirmed their involvement in CP-induced p27 mRNA regulation; while knock-down of the KH-type splicing regulatory protein (KHSRP) further enhanced the sensitivity of MCF7 adenocarcinoma cancer cells to CP treatment. Our results highlight the benefit of specific in vivo mRNA-protein interactome capture to reveal post-transcriptional regulatory networks implicated in cellular drug response and adaptation.
Caspases are key components of apoptotic pathways. Regulation of caspases occurs at several levels, including transcription, proteolytic processing, inhibition of enzymatic function, and protein degradation. In contrast, little is known about the extent of post-transcriptional control of caspases. Here, we describe four conserved RNA-binding proteins (RBPs)—PUF-8, MEX-3, GLD-1, and CGH-1—that sequentially repress the CED-3 caspase in distinct regions of the Caenorhabditis elegans germline. We demonstrate that GLD-1 represses ced-3 mRNA translation via two binding sites in its 3′ untranslated region (UTR), thereby ensuring a dual control of unwanted cell death: at the level of p53/CEP-1 and at the executioner caspase level. Moreover, we identified seven RBPs that regulate human caspase-3 expression and/or activation, including human PUF-8, GLD-1, and CGH-1 homologs PUM1, QKI, and DDX6. Given the presence of unusually long executioner caspase 3′ UTRs in many metazoans, translational control of executioner caspases by RBPs might be a strategy used widely across the animal kingdom to control apoptosis.
The double-stranded RNA-specific editase 1 (RED1/ADAR2) is implicated in the editing of precursor-mRNAs (pre-mRNA) encoding subunits of glutamate receptors (GluRs) in brain. Site-specific deamination of adenosine to inosine alters the codon at the Q/R site in GluR-B rendering the heteromeric receptor impermeable to Ca ions. We cloned human RED1 (hRED1/hADAR2) cDNAs from a brain cDNA library. The human enzyme is 95% identical to the rat homologue. We characterized two alternatively spliced forms that differed by the presence of an Alu-J cassette in the deaminase domain. For the long form containing the Alu cassette, we isolated cDNA clones with an alternative C- terminus and 3'-UTR. An 8.8-kb transcript of hRED1 is most abundant in brain and heart, and lower levels are detected in other tissues. In vitro editing assays with purified recombinant hRED1 containing or lacking the Alu-J cassette revealed that both forms of the protein have the same substrate specificity, but differ in their catalytic activity.
Identifying the interaction partners of noncoding RNAs is essential for elucidating their functions. We have developed an approach, termed microRNA crosslinking and immunoprecipitation (miR-CLIP), using pre-miRNAs modified with psoralen and biotin to capture their targets in cells. Photo-crosslinking and Argonaute 2 immunopurification followed by streptavidin affinity purification of probe-linked RNAs provided selectivity in the capture of targets, which were identified by deep sequencing. miR-CLIP with pre-miR-106a, a miR-17-5p family member, identified hundreds of putative targets in HeLa cells, many carrying conserved sequences complementary to the miRNA seed but also many that were not predicted computationally. miR-106a overexpression experiments confirmed that miR-CLIP captured functional targets, including H19, a long noncoding RNA that is expressed during skeletal muscle cell differentiation. We showed that miR-17-5p family members bind H19 in HeLa cells and myoblasts. During myoblast differentiation, levels of H19, miR-17-5p family members and mRNA targets changed in a manner suggesting that H19 acts as a 'sponge' for these miRNAs.
Homeostatic regulation of ionic currents is of paramount importance during periods of synaptic growth or remodeling. Our previous work has identified the translational repressor Pumilio (Pum) as a regulator of sodium current (I Na) and excitability in Drosophila motoneurons. In this current study, we show that Pum is able to bind directly the mRNA encoding the Drosophila voltage-gated sodium channel paralytic (para). We identify a putative binding site for Pum in the 3′ end of the para open reading frame (ORF). Characterization of the mechanism of action of Pum, using whole-cell patch clamp and real-time reverse transcription-PCR, reveals that the full-length protein is required for translational repression of para mRNA. Additionally, the cofactor Nanos is essential for Pum-dependent para repression, whereas the requirement for Brain Tumor (Brat) is cell type specific. Thus, Pum-dependent regulation of I Na in motoneurons requires both Nanos and Brat, whereas regulation in other neuronal types seemingly requires only Nanos but not Brat. We also show that Pum is able to reduce the level of nanos mRNA and as such identify a potential negative-feedback mechanism to protect neurons from overactivity of Pum. Finally, we show coupling between I Na (para) and I K (Shal) such that Pum-mediated change in para results in a compensatory change in Shal. The identification of para as a direct target of Pum represents the first ion channel to be translationally regulated by this repressor and the location of the binding motif is the first example in an ORF rather than in the canonical 3′-untranslated region of target transcripts.
Post-transcriptional regulation of gene expression plays important roles in diverse cellular processes such as development, neurogenesis and cancer progression. This regulation is often mediated by specific RNA-binding proteins (RBPs) that bind to elements in the untranslated regions (UTRs) of mRNAs and regulate the stability, translation, or localization of the mRNA. Whereas many classical studies explored the cellular role of RBPs with specific mRNA substrates, the recent development of genome-wide analysis tools enables systematic identification of the mRNA substrates of RBPs, and the study of post-transcriptional gene regulation on a global scale. For instance, mRNAs targeted by specific RBPs can be identified by combining biochemical purification of RBPs followed by the analysis of associated RNAs with DNA microarrays. As exemplified for the Pumilio-Fem-3-binding factor (PUF) proteins, for which the mRNA targets have been systematically identified in yeast and Drosophila, the groups of RNAs associated with RBPs often encode functionally or cytotopically related proteins which act in the same biochemical pathway, are parts of a macromolecular complex or localize to the same subcellular structure. Moreover, the global approach enabled the discovery of conserved sequence elements in the RNA which specify protein binding. These studies suggest extensive coordinate regulation of RNAs by RBPs being part of a highly organized and versatile post-transcriptional regulatory system.
Hundreds of RNA-binding proteins (RBPs) control diverse aspects of post-transcriptional gene regulation. To identify novel and unconventional RBPs, we probed high-density protein microarrays with fluorescently labeled RNA and selected 200 proteins that reproducibly interacted with different types of RNA from budding yeast Saccharomyces cerevisiae. Surprisingly, more than half of these proteins represent previously known enzymes, many of them acting in metabolism, providing opportunities to directly connect intermediary metabolism with posttranscriptional gene regulation. We mapped the RNA targets for 13 proteins identified in this screen and found that they were associated with distinct groups of mRNAs, some of them coding for functionally related proteins. We also found that overexpression of the enzyme Map1 negatively affects the expression of experimentally defined mRNA targets. Our results suggest that many proteins may associate with mRNAs and possibly control their fates, providing dense connections between different layers of cellular regulation.
Genome-wide identification of RNAs associated with RNA-binding proteins is crucial for deciphering posttranscriptional regulatory systems. PUMILIO is a member of the evolutionary conserved Puf-family of RNA-binding proteins that repress gene expression posttranscriptionally. We generated transgenic flies expressing affinity-tagged PUMILIO under the control of an ovary-specific promoter, and we purified PUMILIO from whole adult flies and embryos and analyzed associated mRNAs by using DNA microarrays. Distinct sets comprising hundreds of mRNAs were associated with PUMILIO at the two developmental stages. Many of these mRNAs encode functionally related proteins, supporting a model for coordinated regulation of posttranscriptional modules by specific RNA-binding proteins. We identified a characteristic sequence motif in the 3'-untranslated regions of mRNAs associated with PUMILIO, and the sufficiency of this motif for interaction with PUMILIO was confirmed by RNA pull-down experiments with biotinylated synthetic RNAs. The RNA motif strikingly resembles the one previously identified for Puf3p, one of five Saccharomyces cerevisiae Puf proteins; however, proteins encoded by the associated mRNAs in yeast and Drosophila do not appear to be related. The results suggest extensive posttranscriptional regulation by PUMILIO and uncover evolutionary features of this conserved family of RNA-binding proteins.
RNA-binding proteins (RBPs) are essential for post-transcriptional regulation of gene expression. Recent high-throughput screens have dramatically increased the number of experimentally identified RBPs; however, comprehensive identification of RBPs within living organisms is elusive. Here we describe the repertoire of 765 and 594 proteins that reproducibly interact with polyadenylated mRNAs in Saccharomyces cerevisiae and Caenorhabditis elegans, respectively. Furthermore, we report the differential association of mRNA-binding proteins (mRPBs) upon induction of apoptosis in C. elegans L4-stage larvae. Strikingly, most proteins composing mRBPomes, including components of early metabolic pathways and the proteasome, are evolutionarily conserved between yeast and C. elegans. We speculate, on the basis of our evidence that glycolytic enzymes bind distinct glycolytic mRNAs, that enzyme-mRNA interactions relate to an ancient mechanism for post-transcriptional coordination of metabolic pathways that perhaps was established during the transition from the early 'RNA world' to the 'protein world'.
We describe a tandem RNA isolation procedure (TRIP) that enables purification of in vivo formed messenger ribonucleoprotein (mRNP) complexes. The procedure relies on the purification of polyadenylated mRNAs with oligo(dT) beads from cellular extracts, followed by the capture of specific mRNAs with 3'-biotinylated 2'-O-methylated antisense RNA oligonucleotides, which are recovered with streptavidin beads. TRIP was applied to isolate in vivo crosslinked mRNP complexes from yeast, nematodes and human cells for subsequent analysis of RNAs and bound proteins. The method provides a basis for adaptation to other types of polyadenylated RNAs, enabling the comprehensive identification of bound proteins/RNAs, and the investigation of dynamic rearrangement of mRNPs imposed by cellular or environmental cues.
The development of genome-wide analysis tools has prompted global investigation of the gene expression program, revealing highly coordinated control mechanisms that ensure proper spatiotemporal activity of a cell's macromolecular components. With respect to the regulation of RNA transcripts, the concept of RNA regulons, which – by analogy with DNA regulons in bacteria – refers to the coordinated control of functionally related RNA molecules, has emerged as a unifying theory that describes the logic of regulatory RNA-protein interactions in eukaryotes. Hundreds of RNA-binding proteins and small non-coding RNAs, such as microRNAs, bind to distinct elements in target RNAs, thereby exerting specific and concerted control over posttranscriptional events. In this review, we discuss recent reports committed to systematically explore the RNA-protein interaction network and outline some of the principles and recurring features of RNA regulons: the coordination of functionally related mRNAs through RNA-binding proteins or non-coding RNAs, the modular structure of its components, and the dynamic rewiring of RNA-protein interactions upon exposure to internal or external stimuli. We also summarize evidence for robust combinatorial control of mRNAs, which could determine the ultimate fate of each mRNA molecule in a cell. Finally, the compilation and integration of global protein-RNA interaction data has yielded first insights into network structures and provided the hypothesis that RNA regulons may, in part, constitute noise ‘buffers’ to handle stochasticity in cellular transcription.
RNAs encoding subunits of glutamate-gated ion channel receptors are posttranscriptionally modified by RNA editing and alternative splicing. The change in amino acid sequence caused by RNA editing can affect both the kinetics and the permeability of the ion channel receptors to cations. Here, we report the purification of a 90-kDa double-stranded RNA-specific adenosine deaminase from HeLa cell nuclear extract that specifically edits the glutamine codon at position 586 in the pre-mRNA of the glutamate receptor B subunit. Site-specific deamination of an adenosine to an inosine converts the glutamine codon to that of arginine. Recently, a gene encoding a double- stranded-specific editase (RED1) was cloned from a rat brain cDNA library. Antibodies generated against the deaminase domain of its human homolog specifically recognized and inhibited the activity of the 90-kDa enzyme, indicating that we have purified hRED1 the human homolog of rat RED1. This enzyme is distinct from double-stranded RNA-specific adenosine deaminase which we and others have previously purified and cloned.
Post-transcriptional gene regulation is driven by RNA-binding proteins (RBPs). Recent global approaches suggest widespread autoregulation of RBPs through binding to their own mRNA; however, little is known about the regulatory impact and quantitative models remain elusive. By integration of several independent kinetic parameters and abundance data, we modelled autoregulatory feedback loops for six canonical and non-canonical RBPs from the yeast Saccharomyces cerevisiae, namely Hrb1p, Hek2/Khd1p, Ski2p, Npl3p, Pfk2p, and Map1p. By numerically solving ordinary differential equations, we compared non-feedback models with models that considered the RPBs as post-transcriptional activators/repressors of their own expression. While our results highlight a substantial gap between predicted protein output and experimentally determined protein abundances applying a no-feedback model, addition of positive feedback loops are surprisingly versatile and can improve predictions towards experimentally determined protein levels, whereas negative feedbacks are particularly sensitive to cooperativity. Our data suggests that introduction of feedback loops supported by real data can improve models of post-transcriptional gene expression.
We have identified an RNA-specific adenosine deaminase (termed Tad1p/scADAT1) from Saccharomyces cerevisiae that selectively converts adenosine at position 37 of eukaryotic tRNAAla to inosine. The activity of purified recombinant Tad1p depends on the conformation of its tRNA substrate and the enzyme was found to be inactive on all other types of RNA tested. Mutant strains in which the TAD1 gene is disrupted are viable but lack Tad1p enzyme activity and their tRNAAla is not modified at position A37. Transformation of the mutant cells with the TAD1 gene restored enzyme activity. Tad1p has significant sequence similarity with the mammalian editing enzymes which act on specific precursor-mRNAs and on long double-stranded RNA. These findings suggest an evolutionary link between pre-mRNA editing and tRNA modification.
DICER1 syndrome is a cancer predisposition disorder caused by mutations that disrupt the function of DICER1 in miRNA processing. Studying the molecular, cellular and oncogenic effects of these mutations can reveal novel mechanisms that control cell homeostasis and tumor biology. Here, we conduct the first analysis of pathogenic DICER1 syndrome allele from the DICER1 3 UTR. We find that the DICER1 syndrome allele, rs1252940486, abolishes interaction with the PUMILIO RNA binding protein with the DICER1 3 UTR, resulting in the degradation of the DICER1 mRNA by AUF1. This single mutational event leads to diminished DICER1 mRNA and protein levels, and widespread reprogramming of miRNA networks. The in-depth characterization of the rs1252940486 DICER1 allele, reveals important post-transcriptional regulatory events that control DICER1 levels.
Glucose inhibition of gluconeogenic growth suppressor 2 protein (Gis2p) and zinc-finger protein 9 (ZNF9) are conserved yeast and human zinc-finger proteins. The function of yeast Gis2p is unknown, but human ZNF9 has been reported to bind nucleic acids, and mutations in the ZNF9 gene cause the neuromuscular disease myotonic dystrophy type 2. To explore the impact of these proteins on RNA regulation, we undertook a systematic analysis of the RNA targets and of the global implications for gene expression.
RNA-binding proteins play essential roles in the post-transcriptional regulation of gene expression. While hundreds of RNA-binding proteins can be predicted computationally, the recent introduction of proteome-wide approaches has dramatically expanded the repertoire of proteins interacting with RNA. Besides canonical RNA-binding proteins that contain characteristic RNA-binding domains, many proteins that lack such domains but have other well-characterised cellular functions were identified; including metabolic enzymes, heat shock proteins, kinases, as well as transcription factors and chromatin-associated proteins. In the context of these recently published RNA-protein interactome datasets obtained from yeast, nematodes, flies, plants and mammalian cells, we discuss examples for seemingly evolutionary conserved “unconventional” RNA-binding proteins that act in central carbon metabolism, stress response or regulation of transcription.
Fetal hemoglobin, HbF (a2c2), is the main hemoglobin synthesized up to birth, but it subsequently declines and adult hemoglobin, HbA (a2b2), becomes predominant. Several studies have indicated that expression of the HbF subunit c-globin might be regulated post-transcriptionally. This could be confered by ,22-nucleotide long microRNAs that associate with argonaute proteins to specifically target c-globin mRNAs and inhibit protein expression. Indeed, applying immunopurifications, we found that c-globin mRNA was associated with argonaute 2 isolated from reticulocytes that contain low levels of HbF (,1%), whereas association was significantly lower in reticulocytes with high levels of HbF (90%). Comparing microRNA expression in reticulocytes from cord blood and adult blood, we identified several miRNAs that were preferentially expressed in adults, among them miRNA-96. The overexpression of microRNA-96 in human ex vivo erythropoiesis decreased c-globin expression by 50%, whereas the knock-down of endogenous microRNA-96 increased c-globin expression by 20%. Moreover, luciferase reporter assays showed that microRNA-96 negatively regulates expression of c-globin in HEK293 cells, which depends on a seedless but highly complementary target site located within the coding sequence of c-globin. Based on these results we conclude that microRNA-96 directly suppresses c-globin expression and thus contributes to HbF regulation.