Nature Inspired Computing and Engineering Research Group
Nature presents some of the best examples of how to solve complex problems efficiently and effectively.
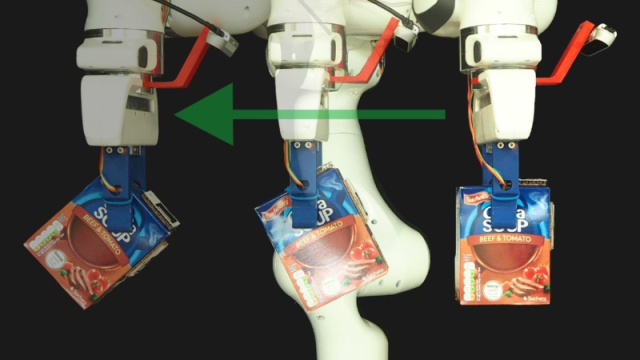
In our group, we advance the foundations of artificial intelligence by drawing inspiration from nature and foundational science disciplines, for example from cognitive sciences, including neuroscience and psychology, and also biological systems, including gene regulatory networks, natural evolution, and physical sciences. We apply our techniques to solve real-world problems in healthcare and sustainability, computer vision and robotics, natural language processing, and other areas.
The Nature Inspired Computing and Engineering group (NICE) is part of the Computer Science Research Centre, together with the Surrey Centre for Cyber Security (SCCS).
Meet the team
Dr Roman Bauer
Head of the Nature Inspired Computing and Engineering Group
Stay connected
Latest tweets
@NICEGroupSurrey