Professor Johnjoe McFadden
About
My qualifications
Previous roles
News
ResearchResearch interests
- Systems Biology
- Mycobacterial genetics
- Pathogenicity of tuberculosis
- Neisserial genetics
- Pathogenicity of meningococcal meningitis
- Quantum Biology - It has recently been recognized that non-trivial aspects of quantum mechanics play a major role in biology:
- See presentations from a workshop on this topic held at the University of Surrey,
- Read review in The Biologist.
Research collaborations
Mycobacterial research
The tubercle bacillus (Mycobacterium tuberculosis) infects approximately one quarter of the world's population and is responsible for three million deaths each year. The related pathogen, Mycobacterium bovis, causes disease in many mammals and is a major cause of economic loss in livestock, particularly cows.
Mycobacterial research within the Microbial Sciences Group is focussed on understanding the pathogenic mechanisms that allow Mycobacterium tuberculosis to cause about three million deaths each year, identification of new drug targets and development of new vaccines to treat the disease.
Most of our work utilises a systems biology modelling approach, particularly applied to metabolism. We constructed the first genome-scale metabolic model of the TB bacillus and used the model to analyse the metabolism of the pathogen both in the laboratory (http://epubs.surrey.ac.uk/184899/) and in infected cells .
The following projects are ongoing:
- Identification of nitrogen source and metabolism of Mycobacterium tuberculosis during intracellular replication. BBSRC-funded.
- Development of recombinant BCG vaccine and complementary diagnostics for TB control in cattle. BBSRC-funded
- BBSRC Investigation of stochastic variations in growth rate as the mechanism of drug tolerance in Mycobacterium tuberculosis. BBSRC-funded.
- Improving BCG vaccination to make it compatible with the skin test. Gates Foundation-funded.
- Defining the metabolic phenotype of intracellular Mycobacterium tuberculosis. (PI, Dany Beste) MRC-funded.
SurreyFBA: Interactive tool for computer simulations of genome scale metabolic networks. (PI, Andrzej Kierzek), BBSRC-funded.
Meningococcal research
Our work has been to investigate virulence mechanisms in Neisseria meningitidis with the aim of developing new vaccines capable of protecting against all strains. Recent successes of our laboratory include constructing a genome-scale metabolic model of the meningococcusand performing an immunoproteomic study of the meningococcus.
Neisseria meningitidis cells as revealed by scanning electron microscopy. The cells tend to occur in pairs and are surrounded by a waxy capsule that protects them from immune attack. Most of the current meningococcal vaccines generate antibodies to the capsule that kills the pathogen. However, antibodies cannot be generated against Group B strains of the meningococcus, which is the most common cause of bacterial meningitis in the UK. Work in our laboratory aims to identify alternative targets of vaccine immunity.
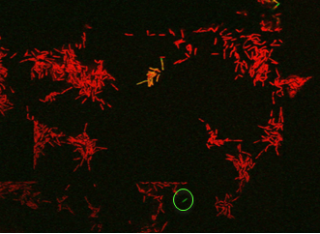
When bacteria, such as Mycobacterium tuberculosis, are treated with an antibiotic a very small fraction survive, despite being genetically identical to the killed population. Persisters can be seen in this video which shows growth of Mycobacterium smegmatis and killing with an antibiotic (4 hours). Killed cells are revealed by their permeability to a fluorescent dye. However, one cell survives and, when the antibiotic is removed (at 14 hours), it is able to grow. The cell is not genetically resistant to the antibiotic as, when antibiotic is added again (at 24 hours), all its descendent cells are killed: it is a persister.
These ‘persisters’ are a major problem in treatment of bacterial infections, particularly in tuberculosis; and how they manage to survive exposure to antibiotics is a mystery that is being activity investigated here at the University of Surrey.
Mycobacterium smegmatis
When bacteria, such as Mycobacterium tuberculosis, are treated with an antibiotic a very small fraction survive, despite being genetically identical to the killed population. Persisters can be seen in this video which shows growth of Mycobacterium smegmatis and killing with an antibiotic (4 hours). Killed cells are revealed by their permeability to a fluorescent dye. However, one cell survives and, when the antibiotic is removed (at 14 hours), it is able to grow. The cell is not genetically resistant to the antibiotic as, when antibiotic is added again (at 24 hours), all its descendent cells are killed: it is a persister.
These ‘persisters’ are a major problem in treatment of bacterial infections, particularly in tuberculosis; and how they manage to survive exposure to antibiotics is a mystery that is being activity investigated here at the University of Surrey.
Research interests
- Systems Biology
- Mycobacterial genetics
- Pathogenicity of tuberculosis
- Neisserial genetics
- Pathogenicity of meningococcal meningitis
- Quantum Biology - It has recently been recognized that non-trivial aspects of quantum mechanics play a major role in biology:
- See presentations from a workshop on this topic held at the University of Surrey,
- Read review in The Biologist.
Research collaborations
Mycobacterial research
The tubercle bacillus (Mycobacterium tuberculosis) infects approximately one quarter of the world's population and is responsible for three million deaths each year. The related pathogen, Mycobacterium bovis, causes disease in many mammals and is a major cause of economic loss in livestock, particularly cows.
Mycobacterial research within the Microbial Sciences Group is focussed on understanding the pathogenic mechanisms that allow Mycobacterium tuberculosis to cause about three million deaths each year, identification of new drug targets and development of new vaccines to treat the disease.
Most of our work utilises a systems biology modelling approach, particularly applied to metabolism. We constructed the first genome-scale metabolic model of the TB bacillus and used the model to analyse the metabolism of the pathogen both in the laboratory (http://epubs.surrey.ac.uk/184899/) and in infected cells .
The following projects are ongoing:
- Identification of nitrogen source and metabolism of Mycobacterium tuberculosis during intracellular replication. BBSRC-funded.
- Development of recombinant BCG vaccine and complementary diagnostics for TB control in cattle. BBSRC-funded
- BBSRC Investigation of stochastic variations in growth rate as the mechanism of drug tolerance in Mycobacterium tuberculosis. BBSRC-funded.
- Improving BCG vaccination to make it compatible with the skin test. Gates Foundation-funded.
- Defining the metabolic phenotype of intracellular Mycobacterium tuberculosis. (PI, Dany Beste) MRC-funded.
SurreyFBA: Interactive tool for computer simulations of genome scale metabolic networks. (PI, Andrzej Kierzek), BBSRC-funded.
Meningococcal research
Our work has been to investigate virulence mechanisms in Neisseria meningitidis with the aim of developing new vaccines capable of protecting against all strains. Recent successes of our laboratory include constructing a genome-scale metabolic model of the meningococcusand performing an immunoproteomic study of the meningococcus.
Neisseria meningitidis cells as revealed by scanning electron microscopy. The cells tend to occur in pairs and are surrounded by a waxy capsule that protects them from immune attack. Most of the current meningococcal vaccines generate antibodies to the capsule that kills the pathogen. However, antibodies cannot be generated against Group B strains of the meningococcus, which is the most common cause of bacterial meningitis in the UK. Work in our laboratory aims to identify alternative targets of vaccine immunity.
When bacteria, such as Mycobacterium tuberculosis, are treated with an antibiotic a very small fraction survive, despite being genetically identical to the killed population. Persisters can be seen in this video which shows growth of Mycobacterium smegmatis and killing with an antibiotic (4 hours). Killed cells are revealed by their permeability to a fluorescent dye. However, one cell survives and, when the antibiotic is removed (at 14 hours), it is able to grow. The cell is not genetically resistant to the antibiotic as, when antibiotic is added again (at 24 hours), all its descendent cells are killed: it is a persister.
These ‘persisters’ are a major problem in treatment of bacterial infections, particularly in tuberculosis; and how they manage to survive exposure to antibiotics is a mystery that is being activity investigated here at the University of Surrey.
Mycobacterium smegmatis
When bacteria, such as Mycobacterium tuberculosis, are treated with an antibiotic a very small fraction survive, despite being genetically identical to the killed population. Persisters can be seen in this video which shows growth of Mycobacterium smegmatis and killing with an antibiotic (4 hours). Killed cells are revealed by their permeability to a fluorescent dye. However, one cell survives and, when the antibiotic is removed (at 14 hours), it is able to grow. The cell is not genetically resistant to the antibiotic as, when antibiotic is added again (at 24 hours), all its descendent cells are killed: it is a persister.
These ‘persisters’ are a major problem in treatment of bacterial infections, particularly in tuberculosis; and how they manage to survive exposure to antibiotics is a mystery that is being activity investigated here at the University of Surrey.
Publications
In 2023, Mycobacterium tuberculosis (Mtb) caused 10.6 million new tuberculosis cases and 1.3 million deaths. The WHO proscribed treatment is not always successful, even when strains were sensitive to the antibiotics.as clinical Mtb populations contain phenotypically tolerant subpopulations, termed persisters. Here a Mtb transposon library was challenged with rifampicin (RIF) and streptomycin (STM) under conditions designed to identify genes that modulate persister frequency. Mutants with reduced survival in RIF were predominantly in genes associated with membrane integrity e.g. arabinogalactan assembly genes cpsA/lytR/Psr, whilst for STM, reduced survival was associated with toxin/antitoxin genes. Some mutations enhanced survival. For RIF these included the methyl citrate cycle genes prpC, prpD and prpR, and the trkA-C K uptake system genes ceoB and Rv2690, and for STM, the resistance associated gene, gidB, and anion-transport genes Rv3679c and Rv3680c. Few genes overlapped the RIF and STM selections, demonstrating that survival mechanisms were antibiotic-specific. Directed deletions of ΔprpD and ΔfadE5 confirmed their predicted enhanced and reduced RIF fitness respectively. The study identified genes that modulate not only persister frequency but also resistance and tolerance, and demonstrates that the mechanisms that produce these phenotypes are diverse and antibiotic-specific.
The biological effects of electromagnetic fields (EMFs) on the central nervous system (CNS) have been widely reported in the literature. Their nature and extent are thought to depend on parameters such as field intensity and frequency. Of these, extremely low-frequency (50 Hz) fields have been reported to influence neuronal firing in CNS regions, including the hippocampus. We applied the loose patch clamp technique to study the effects of 1 mT exposures of such fields over the course of 60 min on cornus ammonis 1 (CA1) pyramidal neuron membranes in coronal hippocampal slices. Such exposure decreased both inward and transient outward currents. Pharmacological blockers of ryanodine receptor (RyR)-dependent Ca release (dantrolene) and endoplasmic reticular Ca store reuptake (SERCA; cyclopiazonic acid) both abrogated these effects. We thus implicate Ca homeostasis in an EMF-induced modulation of neuronal excitability through its regulation of voltage-gated channels.
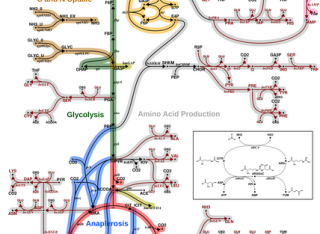
Metabolic flux is the final output of cellular regulation and has been extensively studied for carbon but much less is known about nitrogen, which is another important building block for living organisms. For the tuberculosis pathogen, this is particularly important in informing the development of effective drugs targeting the pathogen's metabolism. Here we performed 13C15N dual isotopic labeling of Mycobacterium bovis BCG steady state cultures, quantified intracellular carbon and nitrogen fluxes and inferred reaction bidirectionalities. This was achieved by model scope extension and refinement, implemented in a multi-atom transition model, within the statistical framework of Bayesian model averaging (BMA). Using BMA-based 13C15N-metabolic flux analysis, we jointly resolve carbon and nitrogen fluxes quantitatively. We provide the first nitrogen flux distributions for amino acid and nucleotide biosynthesis in mycobacteria and establish glutamate as the central node for nitrogen metabolism. We improved resolution of the notoriously elusive anaplerotic node in central carbon metabolism and revealed possible operation modes. Our study provides a powerful and statistically rigorous platform to simultaneously infer carbon and nitrogen metabolism in any biological system.
Persistence has been linked to treatment failure since its discovery over 70 years ago and understanding formation, nature and survival of this key antibiotic refractory subpopulation is crucial to enhancing treatment success and combatting the threat of antimicrobial resistance (AMR). The term ‘persistence’ is often used interchangeably with other terms such as tolerance or dormancy. In this review we focus on ‘antibiotic persistence’ which we broadly define as a feature of a subpopulation of bacterial cells that possesses the non-heritable character of surviving exposure to one or more antibiotics; and persisters as cells that possess this character. We discuss novel molecular mechanisms involved in persister cell formation, as well as environmental factors which can contribute to increased antibiotic persistence in vivo, highlighting recent developments advanced by single-cell studies. We also aim to provide a comprehensive model of persistence, the ‘hunker’ theory which is grounded in intrinsic heterogeneity of bacterial populations and a myriad of ‘hunkering down’ mechanisms which can contribute to antibiotic survival of the persister subpopulation. Finally, we discuss antibiotic persistence as a ‘stepping-stone’ to AMR and stress the urgent need to develop effective anti-persister treatment regimes to treat this highly clinically relevant bacterial sub-population.
The quest to comprehend the nature of consciousness has spurred the development of many theories that seek to explain its underlying mechanisms and account for its neural correlates. In this paper, I compare my own conscious electromagnetic information field (cemi field) theory with integrated information theory (IIT) and global workspace theory (GWT) for their ability to 'carve nature at its joints' in the sense of predicting the entities, structures, states and dynamics that are conventionally recognized as being conscious or nonconscious. I go on to argue that, though the cemi field theory shares features of both integrated information theory and global workspace theory, it is more successful at carving nature at its conventionally accepted joints between conscious and nonconscious systems, and is thereby a more successful theory of consciousness.
Hippocampal pyramidal neuronal activity has been previously studied using conventional patch clamp in isolated cells and brain slices. We here introduce the loose patch clamping study of voltage-activated currents from in situ pyramidal neurons in murine cornus ammonis 1 hippocampal coronal slices. Depolarizing pulses of 15-ms duration elicited early transient inward, followed by transient and prolonged outward currents in the readily identifiable junctional region between the stratum pyramidalis (SP) and oriens (SO) containing pyramidal cell somas and initial segments. These resembled pyramidal cell currents previously recorded using conventional patch clamp. Shortening the depolarizing pulses to >1-2 ms continued to evoke transient currents; hyperpolarizing pulses to varying voltages evoked decays whose time constants could be shortened to
Understanding the rules of life is one of the most important scientific endeavours and has revolutionised both biology and biotechnology. Remarkable advances in observation techniques allow us to investigate a broad range of complex and dynamic biological processes in which living systems could exploit quantum behaviour to enhance and regulate biological functions. Recent evidence suggests that these non-trivial quantum mechanical effects may play a crucial role in maintaining the non-equilibrium state of biomolecular systems. Quantum biology is the study of such quantum aspects of living systems. In this review, we summarise the latest progress in quantum biology, including the areas of enzyme-catalysed reactions, photosynthesis, spin-dependent reactions, DNA, fluorescent proteins, and ion channels. Many of these results are expected to be fundamental building blocks towards understanding the rules of life.
One of the most challenging problems in microbiology is to understand how a small fraction of microbes that resists killing by antibiotics can emerge in a population of genetically identical cells, the phenomenon known as persistence or drug tolerance. Its characteristic signature is the biphasic kill curve, whereby microbes exposed to a bactericidal agent are initially killed very rapidly but then much more slowly. Here we relate this problem to the more general problem of understanding the emergence of distinct growth phenotypes in clonal populations. We address the problem mathematically by adopting the framework of the phenomenon of so-called weak ergodicity breaking, well known in dynamical physical systems, which we extend to the biological context. We show analytically and by direct stochastic simulations that distinct growth phenotypes can emerge as a consequence of slow-down of stochastic fluctuations in the expression of a gene controlling growth rate. In the regime of fast gene transcription, the system is ergodic, the growth rate distribution is unimodal, and accounts for one phenotype only. In contrast, at slow transcription and fast translation, weakly non-ergodic components emerge, the population distribution of growth rates becomes bimodal, and two distinct growth phenotypes are identified. When coupled to the well-established growth rate dependence of antibiotic killing, this model describes the observed fast and slow killing phases, and reproduces much of the phenomenology of bacterial persistence. The model has major implications for efforts to develop control strategies for persistent infections.
Conventional theories of consciousness (ToCs) that assume that the substrate of consciousness is the brain's neuronal matter fail to account for fundamental features of consciousness, such as the binding problem. Field ToC's propose that the substrate of consciousness is the brain's best accounted by some kind of field in the brain. Electromagnetic (EM) ToCs propose that the conscious field is the brain's well-known EM field. EM-ToCs were first proposed only around 20 years ago primarily to account for the experimental discovery that synchronous neuronal firing was the strongest neural correlate of consciousness (NCC). Although EM-ToCs are gaining increasing support, they remain controversial and are often ignored by neurobiologists and philosophers and passed over in most published reviews of consciousness. In this review I examine EM-ToCs against established criteria for distinguishing between ToCs and demonstrate that they outperform all conventional ToCs and provide novel insights into the nature of consciousness as well as a feasible route toward building artificial consciousnesses.
Persistence, the phenomenon whereby a small subpopulation of bacterial cells survive sterilization, prolongs antibiotic treatment and contributes to the development of genetic antimicrobial drug resistance (AMR). In this study we performed single-cell tracking of wild-type and high-persister mutant strains of Escherichia coli to identify factors that correlate with persistence. We found, as expected, persistence correlated with slow growth, but also with small birth size. We investigated intergenerational (mother–daughter) and intragenerational (sister–sister) phenotypic inheritance of growth parameters and discovered the mutant phenotype was associated with lower levels of phenotypic inheritance and identified the gene responsible, the transcription factor ydcI. Targeting pathways involved in persistence could reveal approaches to impeding persistence and the development of AMR.
The use of carbon nanotubes as a gene delivery system has been extensively studied in recent years owing to its potential advantages over viral vectors. To achieve this goal, carbon nanotubes have to be functionalized to become compatible with aqueous media and to bind the genetic material. To establish the best conditions for plasmid DNA binding, we compare the dispersion properties of single-, double- and multi-walled carbon nanotubes (SWCNTs, DWCNTs and MWCNTs, respectively) functionalized with a variety of surfactants by non-covalent attachment. The DNA binding properties of the functionalized carbon nanotubes were studied and compared by electrophoresis. Furthermore, a bilayer functionalization method for DNA binding on SWCNTs was developed that utilized RNA-wrapping to solubilize the nanotubes and cationic polymers as a bridge between nanotubes and DNA.
Carbon nanotubes (CNTs) are at present being considered as potential nanovectors with the ability to deliver therapeutic cargoes into living cells. Previous studies established the ability of CNTs to enter cells and their therapeutic utility, but an appreciation of global intracellular trafficking associated with their cellular distribution has yet to be described. Despite the many aspects of the uptake mechanism of CNTs being studied, only a few studies have investigated internalization and fate of CNTs inside cells in detail. In the present study, intracellular localization and trafficking of RNA-wrapped, oxidized double-walled CNTs (oxDWNT-RNA) is presented. Fixed cells, previously exposed to oxDWNT-RNA, were subjected to immunocytochemical analysis using antibodies specific to proteins implicated in endocytosis; moreover cell compartment markers and pharmacological inhibitory conditions were also employed in this study. Our results revealed that an endocytic pathway is involved in the internalization of oxDWNT-RNA. The nanotubes were found in clathrin-coated vesicles, after which they appear to be sorted in early endosomes, followed by vesicular maturation, become located in lysosomes. Furthermore, we observed co-localization of oxDWNT-RNA with the small GTP-binding protein (Rab 11), involved in their recycling back to the plasma membrane via endosomes from the trans-golgi network.
Mycobacterium tuberculosis infects a third of the world's population. Primary tuberculosis involving active fast bacterial replication is often followed by asymptomatic latent tuberculosis, which is characterised by slow or non-replicating bacteria. Reactivation of the latent infection involving a switch back to active bacterial replication can lead to post-primary transmissible tuberculosis. Mycobacterial mechanisms involved in slow growth or switching growth rate provide rational targets for the development of new drugs against persistent mycobacterial infection. Using chemostat culture to control growth rate, we screened a transposon mutant library by Transposon site hybridization (TraSH) selection to define the genetic requirements for slow and fast growth of Mycobacterium bovis (BCG) and for the requirements of switching growth rate. We identified 84 genes that are exclusively required for slow growth (69 hours doubling time) and 256 genes required for switching from slow to fast growth. To validate these findings we performed experiments using individual M. tuberculosis and M. bovis BCG knock out mutants. We have demonstrated that growth rate control is a carefully orchestrated process which requires a distinct set of genes encoding several virulence determinants, gene regulators, and metabolic enzymes. The mce1 locus appears to be a component of the switch to slow growth rate, which is consistent with the proposed role in virulence of M. tuberculosis. These results suggest novel perspectives for unravelling the mechanisms involved in the switch between acute and persistent TB infections and provide a means to study aspects of this important phenomenon in vitro.
The co-catabolism of multiple host-derived carbon substrates is required by Mycobacterium tuberculosis (Mtb) to successfully sustain a tuberculosis infection. However, the metabolic plasticity of this pathogen and the complexity of the metabolic networks present a major obstacle in identifying those nodes most amenable to therapeutic interventions. It is therefore critical that we define the metabolic phenotypes of Mtb in different conditions. We applied metabolic flux analysis using stable isotopes and lipid fingerprinting to investigate the metabolic network of Mtb growing slowly in our steady-state chemostat system. We demonstrate that Mtb efficiently co-metabolises either cholesterol or glycerol, in combination with two-carbon generating substrates without any compartmentalisation of metabolism. We discovered that partitioning of flux between the TCA cycle and the glyoxylate shunt combined with a reversible methyl citrate cycle is the critical metabolic nodes which underlie the nutritional flexibility of Mtb. These findings provide novel insights into the metabolic architecture that affords adaptability of bacteria to divergent carbon substrates and expand our fundamental knowledge about the methyl citrate cycle and the glyoxylate shunt.
Whenever a genetically homogenous population of bacterial cells is exposed to antibiotics, a tiny fraction of cells survives the treatment,the phenomenon known as bacterial persistence [G.L. Hobby et al., Exp. Biol. Med. 50, 281–285 (1942); J. Bigger, The Lancet 244, 497– 500 (1944)]. Despite its biomedical relevance, the origin of the phenomenon is still unknown, and as a rare, phenotypically resistant subpopulation, persisters are notoriously hard to study and define. Using computerized tracking we show that persisters are small at birth and slowly replicating. We also determine that the highpersister mutant strain of Escherichia coli, HipQ, is associated with the phenotype of reduced phenotypic inheritance (RPI). We identify the gene responsible for RPI, ydcI, which encodes a transcription factor, and propose a mechanism whereby loss of phenotypic inheritance causes increased frequency of persisters. These results provide insight into the generation and maintenance of phenotypic variation and provide potential targets for the development of therapeutic strategies that tackle persistence in bacterial infections.
The tuberculin purified protein derivative (PPD) is a widely used diagnostic antigen for tuberculosis, however it is poorly defined. Most mycobacterial proteins are extensively denatured by the procedure employed in its preparation, which explains previous difficulties in identifying constituents from PPD to characterize their behaviour in B- and T-cell reactions. We here described a proteomics-based characterization of PPD from several different sources by LC-MS/MS, which combines the solute separation power of HPLC, with the detection power of a mass spectrometer. The technique is able to identify proteins from complex mixtures of peptide fragments. A total of 171 different proteins were identified among the four PPD samples (two bovine PPD and two avium PPD) from Brazil and UK. The majority of the proteins were cytoplasmic (77.9%) and involved in intermediary metabolism and respiration (24.25%) but there was a preponderance of proteins involved in lipid metabolism. We identified a group of 21 proteins that are present in both bovine PPD but were not detected in avium PPD preparation. In addition, four proteins found in bovine PPD are absent in Mycobacterium bovis BCG vaccine strain. This study provides a better understanding of the tuberculin PPD components leading to the identification of additional antigens useful as reagents for specific diagnosis of tuberculosis.
The cause of sarcoidosis is unknown. However, the histological similarity between the disorder and tuberculosis suggests that mycobacteria might contribute to the pathogenesis of sarcoidosis. We have used the polymerase chain reaction (PCR) to detect mycobacterial DNA in clinical samples from patients with sarcoidosis. 104 patients were included in the study (62 referred for possible tuberculosis and 20 for possible sarcoidosis, and 22 control patients who had undergone bronchoscopy for other reasons). Bronchoalveolar lavage samples, bronchial washings, and tissue specimens (1 from each patient) underwent assay by PCR as well as bacteriological, histological, and cytological examination. We used two PCR reactions: in the first the complex-specific insertion sequence IS986/IS6110 was used to specifically detect DNA from Mycobacterium tuberculosis complex bacteria; in the second, conserved sequences of the mycobacterial groEL gene were used to detect DNA from mycobacteria other than M tuberculosis. The PCR was more sensitive than culture for diagnosis of tuberculosis. However, the false-positive PCR rate for M tuberculosis was 9%. M tuberculosis DNA was found in half the sarcoidosis patients, and non-tuberculosis mycobacterial DNA in a further 20%. The findings that a significant proportion of the sarcoidosis patients in this study have mycobacteria in their lungs and that most of these mycobacteria belong to M tuberculosis complex suggest an aetiological role for mycobacteria in sarcoidosis.
Today, hyperthermophilic ('superheat-loving') bacteria and archaea are found within high-temperature environments, representing the upper temperature border of life. They grow optimally above 80°C and exhibit an upper temperature border of growth up to 113°C. Members of the genera, Pyrodictium and Pyrolobus, survive at least 1 h of autoclaving. In their basically anaerobic environments, hyperthermophiles (HT) gain energy by inorganic redox reactions employing compounds like molecular hydrogen, carbon dioxide, sulphur and ferric and ferrous iron. Based on their growth requirements, HT could have existed already on the early Earth about 3.9 Gyr ago. In agreement, within the phylogenetic tree of life, they occupy all the short deep branches closest to the root. The earliest archaeal phylogenetic lineage is represented by the extremely tiny members of the novel kingdom of Nanoarchaeota, which thrive in submarine hot vents. HT are very tough survivors, even in deep-freezing at −140°C. Therefore, during impact ejecta, they could have been successfully transferred to other planets and moons through the coldness of space.
An insertion sequence (IS901), found in pathogenic strains of Mycobacterium avium, but absent in M. avium complex isolates from patients with acquired immune deficiency syndrome (AIDS), has been isolated and sequenced. This insertion element has a nucleotide sequence of 1472 bp, with one open reading frame (0RF1), which codes for a protein of 401 amino acids. The amino acid sequence, terminal ends and target site of IS901 are similar to those of IS900, present in Mycobacterium paratuberculosis. However, the DNA sequences of these two IS elements exhibit only 60% homology, compared to a DNA homology of 98% between their respective hosts. IS901, like IS900, appears to belong to a family of related insertion elements present in actinomycetes and other bacteria. M. avium strains containing IS900 were found to be more virulent in mice than closely related strains lacking IS901. IS901 may be a useful tool for the study of the genetics of virulence in the M. avium complex and for obtaining stable integration of foreign genes into mycobacteria.
Objectives: To determine the occurrence and distribution of IS 1110 in a sample of clinical isolates; to investigate the polymorphism detected with IS 1110-derived probes, and the stability of such patterns; and to evaluate IS 1110-based probes, in comparison with other methods, for differentiation of Mycobacterium avium isolates. Design: Fifty M. avium complex strains used for evaluation of the IS 1110 probe originated from the Memorial Sloan-Kettering Cancer Center, New York. Results: IS 1110 hybridizes to a highly polymorphic element in most M. avium strains. Most banding patterns were found to be unique, but four groups of identical strains were identified. One group, from non-AIDS subjects, was associated with colonization rather than dissemination or invasion. Combining pMB22 and IS 1110 typing yielded higher discrimination than either probe alone. Comparison of IS 1110 and pMB22 polymorphisms with multilocus enzyme electrophoresis indicated that the three methods were essentially independent. Conclusions: IS 1110 provides a convenient method for differentiating M. avium isolates for epidemiologic purposes.
Summary Gene replacement by homologous recombination is a powerful tool for fundamental studies of gene function, as well as allowing specific attenuation of pathogens, but has proved difficult to achieve for Mycobacterium tuberculosis. We have used a plasmid‐based test system to demonstrate the occurrence of homologous recombination in the tuberculosis vaccine strain Mycobacterium bovis BCG, and we have successfully replaced a target gene in BCG by homologous recombination, using a shuttle plasmid. Specific inactivation of selected genes will facilitate study of virulence factors and drug resistance as well as allowing rational attenuation of M. tuberculosis for the production of new vaccines.
Nitrogen metabolism of Mycobacterium tuberculosis(Mtb) is crucial for the survival of this important pathogen in its primary human host cell, the macrophage, but little is known about the source(s) and their assimilation within this intracellular niche. Here, we have developed 15N-flux spectral ratio analysis(15N-FSRA) to explore Mtb’s nitrogen metabolism; we demonstrate that intracellular Mtb has access to multiple amino acids in the macrophage, including glutamate, glutamine, aspartate, alanine, glycine,and valine; and we identify glutamine as the pre-dominant nitrogen donor. Each nitrogen source is uniquely assimilated into specific amino acid pools,indicating compartmentalized metabolism during intracellular growth. We have discovered that serine is not available to intracellular Mtb, and we show that a serine auxotroph is attenuated in macrophages. This work provides a systems-based tool for exploring the nitrogen metabolism of intracellular pathogens and highlights the enzyme phosphoserine transaminase as an attractive target for the development of novel anti-tuberculosis therapies.
The role of mycobacteria, specifically Mycobacterium paratuberculosis, in Crohn's disease has aroused considerable controversy for many years. Using the ultra sensitive polymerase chain reaction some studies have reported detection of M paratuberculosis DNA in as many as 65% of Crohn's disease patients but also in patients without disease. Other studies have been negative for both groups. We therefore designed a double blind control trial to investigate the presence of mycobacterial DNA in age, sex, and tissue matched paraffin wax embedded tissues from 31 Crohn's disease tissues, 20 diseased gut control tissues, and 10 ulcerative colitis tissues. The specimens were coded and analysed blind with three separate polymerase chain reactions (PCR) based on DNA sequences specific for M paratuberculosis (IS900), M avium (RFLP type A/1) (IS901), and the Mycobacterium genus (65 kDa gene, TB600). The number of granulomata and presence of acid fast bacilli in each Crohn's disease tissue was also investigated. The sensitivity of the system was determined using similarly prepared gut tissue from an animal infected with M paratuberculosis. Four of 31 Crohn's disease tissues and none of the 30 control and ulcerative colitis derived tissues amplified M paratuberculosis DNA. Crohn's disease tissues containing granulomata were significantly more likely to amplify M paratuberculosis specific DNA on PCR than the non-Crohn's disease tissues (p = 0.02). All the positive Crohn's disease tissues contained granulomata, and none contained acid fast bacilli. Equivalent numbers of Crohn's and non-Crohn's disease tissues amplified the region of the 65 kD gene on PCR for non-specific mycobacterial DNA (11/31 and 9/30 respectively). No sections produced an amplified product with the IS901 PCR. These results suggest that few Crohn's disease gut biopsy sections contain M paratuberculosis DNA in association with granulomata. The absence of such DNA in any control and ulcerative colitic tissue strengthens the case for it having a specific association, which may be pathogenic, with Crohn's disease in this minority of patients.
Leptospirosis is a neglected disease of man and animals that affects nearly half a million people annually and causes considerable economic losses. Current human vaccines are inactivated whole-cell preparations (bacterins) of Leptospira spp. that provide strong homologous protection yet fail to induce a cross-protective immune response. Yearly boosters are required, and serious side-effects are frequently reported so the vaccine is licensed for use in humans in only a handful of countries. Novel universal vaccines require identification of conserved surface-exposed epitopes of leptospiral antigens. Outer membrane β-barrel proteins (βb-OMPs) meet these requirements and have been successfully used as vaccines for other diseases. We report the evaluation of 22 constructs containing protein fragments from 33 leptospiral βb-OMPs, previously identified by reverse and structural vaccinology and cell-surface immunoprecipitation. Three-dimensional structures for each leptospiral βb-OMP were predicted by I-TASSER. The surface-exposed epitopes were predicted using NetMHCII 2.2 and BepiPred 2.0. Recombinant constructs containing regions from one or more βb-OMPs were cloned and expressed in Escherichia coli . IMAC-purified recombinant proteins were adsorbed to an aluminium hydroxide adjuvant to produce the vaccine formulations. Hamsters (4-6 weeks old) were vaccinated with 2 doses containing 50 – 125 μg of recombinant protein, with a 14-day interval between doses. Immunoprotection was evaluated in the hamster model of leptospirosis against a homologous challenge (10 – 20× ED 50 ) with L. interrogans serogroup Icterohaemorrhagiae serovar Copenhageni strain Fiocruz L1-130. Of the vaccine formulations, 20/22 were immunogenic and induced significant humoral immune responses (IgG) prior to challenge. Four constructs induced significant protection (100%, P< 0.001) and sterilizing immunity in two independent experiments, however, this was not reproducible in subsequent evaluations (0 – 33.3% protection, P > 0.05). The lack of reproducibility seen in these challenge experiments and in other reports in the literature, together with the lack of immune correlates and commercially available reagents to characterize the immune response, suggest that the hamster may not be the ideal model for evaluation of leptospirosis vaccines and highlight the need for evaluation of alternative models, such as the mouse.
BACKGROUND: Neisseria meningitidis is an important human commensal and pathogen that causes several thousand deaths each year, mostly in young children. How the pathogen replicates and causes disease in the host is largely unknown, particularly the role of metabolism in colonization and disease. Completed genome sequences are available for several strains but our understanding of how these data relate to phenotype remains limited. RESULTS: To investigate the metabolism of N. meningitidis we generated and selected a representative Tn5 library on rich medium, a minimal defined medium and in human serum to identify genes essential for growth under these conditions. To relate these data to a systems-wide understanding of the pathogen's biology we constructed a genome-scale metabolic network: Nmb_iTM560. This model was able to distinguish essential and non-essential genes as predicted by the global mutagenesis. These essentiality data, the library and the Nmb_iTM560 model are powerful and widely applicable resources for the study of meningococcal metabolism and physiology. We demonstrate the utility of these resources by predicting and demonstrating metabolic requirements on minimal medium such as a requirement for PEP carboxylase, and by describing the nutritional and biochemical status of N. meningitidis when grown in serum, including a requirement for both the synthesis and transport of amino acids. CONCLUSIONS: This study describes the application of a genome scale transposon library combined with an experimentally validated genome-scale metabolic network of N. meningitidis to identify essential genes and provide novel insight to the pathogen's metabolism both in vitro and during infection.
Despite decades of research, many aspects of the biology of Mycobacterium tuberculosis remain unclear, and this is reflected in the antiquated tools available to treat and prevent tuberculosis and consequently this disease remains a serious public health problem. Important discoveries linking the metabolism of M. tuberculosis and pathogenesis has renewed interest in this area of research. Previous experimental studies were limited to the analysis of individual genes or enzymes, whereas recent advances in computational systems biology and high-throughput experimental technologies now allows metabolism to be studied on a genome scale. In the present article, we discuss the progress being made in applying system-level approaches to study the metabolism of this important pathogen.
Background BCG is the most widely used vaccine of all time and remains the only licensed vaccine for use against tuberculosis in humans. BCG also protects other species such as cattle against tuberculosis, but due to its incompatibility with current tuberculin testing regimens remains unlicensed. BCG’s efficacy relates to its ability to persist in the host for weeks, months or even years after vaccination. It is unclear to what degree this ability to resist the host’s immune system is maintained by a dynamic interaction between the vaccine strain and its host as is the case for pathogenic mycobacteria. Results To investigate this question, we constructed transposon mutant libraries in both BCG Pasteur and BCG Danish strains and inoculated them into bovine lymph nodes. Cattle are well suited to such an assay, as they are naturally susceptible to tuberculosis and are one of the few animal species for which a BCG vaccination program has been proposed. After three weeks, the BCG were recovered and the input and output libraries compared to identify mutants with in vivo fitness defects. Less than 10% of the mutated genes were identified as affecting in vivo fitness, they included genes encoding known mycobacterial virulence functions such as mycobactin synthesis, sugar transport, reductive sulphate assimilation, PDIM synthesis and cholesterol metabolism. Many other attenuating genes had not previously been recognised as having a virulence phenotype. To test these genes, we generated and characterised three knockout mutants that were predicted by transposon mutagenesis to be attenuating in vivo: pyruvate carboxylase, a hypothetical protein (BCG_1063), and a putative cyclopropane-fatty-acyl-phospholipid synthase. The knockout strains survived as well as wild type during in vitro culture and in bovine macrophages, yet demonstrated marked attenuation during passage in bovine lymph nodes confirming that they were indeed involved in persistence of BCG in the host. Conclusion These data show that BCG is far from passive during its interaction with the host, rather it continues to employ its remaining virulence factors, to interact with the host’s innate immune system to allow it to persist, a property that is important for its protective efficacy.
The genetic relationships between various serotypes and serogroups of meningococcal strains were investigated by restriction fragment-length polymorphism (RFLP) analysis using a number of random DNA probes and a probe containing a truncated copy of the meningococcal insertion sequence IS1106. The data were used to estimate genetic distance between all pairs of strains and to construct phylogenetic trees for meningococcal strains. B15: P1.16R strains isolated from cases of systemic meningococcal disease in two health districts with a high incidence of disease were clonal in contrast to similar strains from cases occurring in other parts of the UK. Strains from these areas, which contain a similar genomic deletion, were found to be derived from two distinct lineages within the B15: P1.16R phylogenetic group. RFLP data demonstrated that present serological typing systems for the meningoccus do not necessarily reflect true genetic relationships.
Occam's razor-the principle of simplicity-has recently been attacked as a cultural bias without rational foundation. Increasingly, belief in pseudoscience and mysticism is growing. I argue that inclusion of Occam's razor is an essential factor that distinguishes science from superstition and pseudoscience. I also describe how the razor is embedded in Bayesian inference and argue that science is primarily the means to discover the simplest descriptions of our world.
The book starts with a general introduction into the relevance of systems biology for understanding tuberculosis.
Meningococcal disease is normally suspected on clinical grounds but confirmed by isolation of Neisseria meningitidis from blood or cerebrospinal fluid (CSF), or by detection of gram-negative diplococci in CSF. After parenteral antibiotics are started the isolation rate of meningococci from blood cultures drops from 50% to less than 5% and the chances of CSF being positive by culture or microscopy are also reduced. We used the polymerase chain reaction (PCR) in a blinded study to detect meningococcal DNA in 54 CSF samples from patients with meningococcal disease or from controls. The PCR primers were specific for the meningococcal insertion sequence IS1106. The sensitivity and specificity of this PCR for diagnosis of meningococcal meningitis were both 91%. Sensitivity was not affected by prior antibiotic treatment. The IS1106 PCR is a rapid and sensitive test for confirmation of the diagnosis of meningococcal meningitis.
The Mycobacterium tuberculosis‐specific insertion sequence IS6110/986 has been widely used as a probe because of the multiple polymorphism observed among different strains. To investigate transposition of IS6110, a series of artificially constructed composite transposons containing IS6110 and a kanamycin resistance marker were constructed. The composite transposons were inserted into a conditionally replicating, thermosensitive, Escherichia coli–mycobacterial shuttle vector and introduced into M. smegmatis mc2155. Lawns of transformants were grown at the permissive temperature on kanamycin‐supplemented agar and subsequently prevented from further growth by shifting to the non‐permissive temperature. Under normal atmospheric conditions, kanamycin‐resistant papillae appeared after only about 5–6 weeks of incubation. However, these events were not associated with transposon mobilization. In contrast, lawns that were exposed to a 48 h microaerobic shock generated kanamycin‐resistant papillae after only 6–14 days. These events were generated by conservative transposition of the IS6110 composite transposon into the M. smegmatis chromosome, with loss of the shuttle vector. In common with other IS3 family elements, transposition of IS6110 is thought to be controlled by translational frameshifting. However, we were unable to detect any significant frameshifting within the putative frameshifting site of IS6110, and the level of frameshifting was not affected by microaerobic incubation. The finding that transposition of IS6110 is stimulated by incubation at reduced oxygen tensions may be relevant to transposition of IS6110 in M. tuberculosis harboured within TB lesions.
Antibiotic persistence is a phenomenon observed when genetically susceptible cells survive long-term exposure to antibiotics. These 'persisters' are an intrinsic component of bacterial populations and stem from phenotypic heterogeneity. Persistence to antibiotics is a concern for public health globally, as it increases treatment duration and can contribute to treatment failure. Furthermore, there is a growing array of evidence that persistence is a 'stepping-stone' for the development of genetic antimicrobial resistance. Urinary tract infections (UTIs) are a major contributor to antibiotic consumption worldwide, and are known to be both persistent (i.e. affecting the host for a prolonged period) and recurring. Currently, in clinical settings, routine laboratory screening of pathogenic isolates does not determine the presence or the frequency of persister cells. Furthermore, the majority of research undertaken on antibiotic persistence has been done on lab-adapted bacterial strains. In the study presented here, we characterized antibiotic persisters in a panel of clinical uropathogenic Escherichia coli isolates collected from hospitals in the UK and Australia. We found that a urine-pH mimicking environment not only induces higher levels of antibiotic persistence to meropenem and colistin than standard laboratory growth conditions, but also results in rapid development of transient colistin resistance, regardless of the genetic resistance profile of the isolate. Furthermore, we provide evidence for the presence of multiple virulence factors involved in stress resistance and biofilm formation in the genomes of these isolates, whose activities have been previously shown to contribute to the formation of persister cells.
Approximately every fourth person in the world currently dies of cancer. Although many efficient anticancer drugs have been developed over the last 60 years or more, most therapeutic approaches still lack specificity for their intended site of action in the body, resulting in reduced effectiveness and severe side effects. The emerging field of nanomedicine provides a whole range of materials and techniques to develop customizable drug delivery vehicles that assist the targeting of therapeutic agents to the desired site of action. Amongst these, carbon nanotubes have emerged as promising candidates, being capable of penetrating mammalian cell membranes and allowing for the attachment of high loads of drugs and targeting agents on their surface or the inner cavity. This chapter will discuss the principles of targeted, anticancer chemotherapies and introduce carbon nanotubes as novel tools for vector-based, targeted drug delivery.
Tuberculosis (TB) is a devastating infectious disease that kills over a million people every year. There is an increasing burden of multi drug resistance (MDR) and extensively drug resistance (XDR) TB. New and improved therapies are urgently needed to overcome the limitations of current treatment. The causative agent, Mycobacterium tuberculosis (Mtb) is one of the most successful pathogens that can manipulate host cell environment for adaptation, evading immune defences, virulence, and pathogenesis of TB infection. Host-pathogen interaction is important to establish infection and it involves a complex set of processes. Metabolic cross talk between the host and pathogen is a facet of TB infection and has been an important topic of research where there is growing interest in developing therapies and drugs that target these interactions and metabolism of the pathogen in the host. Mtb scavenges multiple nutrient sources from the host and has adapted its metabolism to survive in the intracellular niche. Advancements in systems-based omic technologies have been successful to unravel host-pathogen interactions in TB. In this review we discuss the application and usefulness of omics in TB research that provides promising interventions for developing anti-TB therapies.
Several theories of consciousness first described about a decade ago, including the conscious electromagnetic information (CEMI) field theory, claimed that the substrate of consciousness is the brain's electromagnetic (EM) field. These theories were prompted by the observation, in many diverse systems, that synchronous neuronal firing, which generates coherent EM fields, was a strong correlate of attention, awareness, and consciousness. However, when these theories were first described there was no direct evidence that synchronous firing was actually functional, rather than an epiphenomenon of brain function. Additionally, any EM field-based consciousness would be a 'ghost in the machine' unless the brain's endogenous EM field is also able to influence neuron firing. Once again, when these theories were first described, there was only indirect evidence that the brain's EM field influenced neuron firing patterns in the brain. In this paper I describe recent experimental evidence which demonstrate that synchronous neuronal firing does indeed have a functional role in the brain; and also that the brain's endogenous EM field is involved in recruiting neurons to synchronously firing networks. The new data point to a new and unappreciated form of neural communication in the brain that is likely to have significance for all theories of consciousness. I describe an extension of the CEMI field theory that incorporates these recent experimental findings and integrates the theory with the 'communication through coherence' hypothesis.
The use of non-viral vectors as delivery systems in gene therapy has been extensively studied recently owing to their advantages over viral vectors. Here, we propose a new gene delivery system based on the use of RNA-wrapped single-walled carbon nanotubes (SWCNTs) complexed with the cationic protein, protamine and the drug chloroquine. Protamine was selected as a cationic protein acting as bridge between negatively charged RNA-wrapped SWCNTs and plasmid DNA. Protamine also contains a nuclear localization signal which enhances the expression of the transfected gene. The drug chloroquine, a lysosomotropic compound which has been reported to increase the transfection efficiency, was attached to RNA-wrapped SWNTs by ionic interactions. The simultaneous delivery of the drug chloroquine with plasmid DNA clearly showed an enhanced gene delivery and expression. The levels of gene expression were quantified using the luciferase reporter gene as model. Optimal conditions for transfection and gene expression were obtained and cytoxicity of the carbon nanotube complexes measured. The optimal complexes were shown to efficiently deliver plasmid DNA for efficient gene expression and may thereby be useful as gene delivery systems for gene therapy. Copyright © 2012 American Scientific Publishers.
A key aspect of consciousness is that it represents bound or integrated information, prompting an increasing conviction that the physical substrate of consciousness must be capable of encoding integrated information in the brain. However, as Ralph Landauer insisted, 'information is physical' so integrated information must be physically integrated. I argue here that nearly all examples of so-called 'integrated information', including neuronal information processing and conventional computing, are only temporally integrated in the sense that outputs are correlated with multiple inputs: the information integration is implemented in time, rather than space, and thereby cannot correspond to physically integrated information. I point out that only energy fields are capable of integrating information in space. I describe the conscious electromagnetic information (cemi) field theory which has proposed that consciousness is physically integrated, and causally active, information encoded in the brain's global electromagnetic (EM) field. I here extend the theory to argue that consciousness implements algorithms in space, rather than time, within the brain's EM field. I describe how the cemi field theory accounts for most observed features of consciousness and describe recent experimental support for the theory. I also describe several untested predictions of the theory and discuss its implications for the design of artificial consciousness. The cemi field theory proposes a scientific dualism that is rooted in the difference between matter and energy, rather than matter and spirit.
The development of diagnostic tests which can readily differentiate between vaccinated and tuberculosis-infected individuals is crucial for the wider utilization of bacillus Calmette-Guérin (BCG) as vaccine in humans and animals. BCG_0092 is an antigen that elicits specific delayed type hypersensitivity reactions similar in size and morphological aspects to that elicited by purified protein derivative, in both animals and humans infected with the tubercle bacilli. We carried out bioinformatics analyses of the BCG_0092 and designed a diagnostic test by using the predicted MHC class I epitopes. In addition, we performed a knockout of this gene by homologous recombination in the BCG vaccine strain to allow differentiation of vaccinated from infected individuals. For that, the flanking sequences of the target gene (BCG_0092)were cloned into a suicide vector. Spontaneous double crossovers, which result in wild type revertants or knockouts were selected using SacB. BCG_0092 is present only in members of the Mycobacterium tuberculosis complex. Eight predicted MHC class I epitopes with potential for immunological diagnosis were defined, allowing the design of a specific diagnostic test. The strategy used to delete the (BCG_0092) gene from BCG was successful. The knockout genotype was confirmed by PCR and by Southern blot. The mutant BCG strain has the potential of inducing protection against tuberculosis without interfering with the diagnostic test based on the use of selected epitopes from BCG_0092.
The expression of many antigens, stimulatory molecules, or even metabolic pathways in mycobacteria such asMycobacterium bovisBCG orM. smegmatiswas made possible through the development of shuttle vectors, and several recombinant vaccines have been constructed. However, gene expression in any of these systems relied mostly on the selection of natural promoters expected to provide the required level of expression by trial and error. To establish a systematic selection of promoters with a range of strengths, we generated a library of mutagenized promoters through error-prone PCR of the strong PL5promoter, originally from mycobacteriophage L5. These promoters were cloned upstream of the enhanced green fluorescent protein reporter gene, and recombinantM. smegmatisbacteria exhibiting a wide range of fluorescence levels were identified. A set of promoters was selected and identified as having high (pJK-F8), intermediate (pJK-B7, pJK-E6, pJK-D6), or low (pJK-C1) promoter strengths in bothM. smegmatisandM. bovisBCG. The sequencing of the promoter region demonstrated that it was extensively modified (6 to 11%) in all of the plasmids selected. To test the functionality of the system, two different expression vectors were demonstrated to allow corresponding expression levels of theSchistosoma mansoniantigen Sm29 in BCG. The approach used here can be used to adjust expression levels for synthetic and/or systems biology studies or for vaccine development to maximize the immune response.
An experimental system of Mycobacterium tuberculosis growth in a carbon-limited chemostat has been established by the use of Mycobacterium bovis BCG as a model organism. For this model, carbon-limited chemostats with low concentrations of glycerol were used to simulate possible growth rates during different stages of tuberculosis. A doubling time of 23 h (D 0.03 h 1) was adopted to represent cells during the acute phase of infection, whereas a lower dilution rate equivalent to a doubling time of 69 h (D 0.01 h 1) was used to model mycobacterial persistence. This chemostat model allowed the specific response of the mycobacterial cell to carbon limitation at different growth rates to be elucidated. The macromolecular (RNA, DNA, carbohydrate, and lipid) and elemental (C, H, and N) compositions of the biomass were determined for steady-state cultures, revealing that carbohydrates and lipids comprised more than half of the dry mass of the BCG cell, with only a quarter of the dry weight consisting of protein and RNA. Consistent with studies of other bacteria, the specific growth rate impacts on the macromolecular content of BCG and the proportions of lipid, RNA, and protein increased significantly with the growth rate. The correlation of RNA content with the growth rate indicates that ribosome production in carbon-limited M. bovis BCG cells is subject to growth rate-dependent control. The results also clearly show that the proportion of lipids in the mycobacterial cell is very sensitive to changes in the growth rate, probably reflecting changes in the amounts of storage lipids. Finally, this study demonstrates the utility of the chemostat model of mycobacterial growth for functional genomic, physiology, and systems biology studies.
A field isolate, 3bla, of the wheat take-all fungus Gaeumannomyces graminis var. tritici was previously shown to be infected with three serologically unrelated viruses, A, B, and C. It is shown that virus B can be separated into two distinct strains, designated B1 and B2. All four viruses, A, B1, B2 and C, were faithfully transmitted into conidia. However, six out of eight single ascospore cultures derived from 3b1a single conidial cultures were shown to be virus-free. The remaining two single ascospore cultures each contained only one virus, which appeared to be the same in each culture. This virus was serological indistinguishable from virus B1, but had dsRNA components of mol. wt. lower than those of virus B1 and had only the smaller of the two capsid polypepide species of virus B1. No DNA provirus molecules homologous to viruses B1, B2 or C could be detected in two of the virus-free ascospore cultures or in one of the virus-infected ascospore cultures. Very low concentrations of virus particles were detected in hyphal tip isolates of G. graminis . After prolonged storage and subculturing of these isolates, the concentration of virus particles had increased to the level of the parent culture from which the hyphal tip isolates were derived.
Bovine tuberculosis (BtB) caused by Mycobacterium bovis remains a major problem in both the developed and developing countries. control of BtB in the UK is carried out by test and slaughter of infected animals, based primarily on the tuberculin skin test (ppD). Vaccination with the attenuated strain of the M. bovis pathogen, BcG, is not used to control bovine tuberculosis in cattle at present, due to its variable efficacy and because it interferes with the PPD test. Diagnostic tests capable of Differentiating Infected from Vaccinated Animals (DIVA) have been developed that detect immune responses to M. bovis antigens absent in BCG; but these are too expensive and insufficiently sensitive to be used for BtB control worldwide. to address these problems we aimed to generate a synergistic vaccine and diagnostic approach that would permit the vaccination of cattle without interfering with the conventional ppD-based surveillance. the approach was to widen the pool of M. bovis antigens that could be used as DiVA targets, by identifying antigenic proteins that could be deleted from BcG without affecting the persistence and protective efficacy of the vaccine in cattle. Using transposon mutagenesis we identified genes that were essential and those that were non-essential for persistence in bovine lymph nodes. We then inactivated selected immunogenic, but non-essential genes in BcG Danish to create a diagnostic-compatible triple knock-out ΔBCG TK strain. The protective efficacy of the ΔBcG tK was tested in guinea pigs experimentally infected with M. bovis by aerosol and found to be equivalent to wild-type BcG. A complementary diagnostic skin test was developed with the antigenic proteins encoded by the deleted genes which did not cross-react in vaccinated or in uninfected guinea pigs. this study demonstrates the functionality of a new and improved BcG strain which retains its protective efficacy but is diagnostically compatible with a novel DIVA skin test that could be implemented in control programmes.
Despite decades of research, many aspects of the biology of Mycobacterium tuberculosis remain unclear, and this is reflected in the antiquated tools available to treat and prevent tuberculosis and consequently this disease remains a serious public health problem. Important discoveries linking the metabolism of M. tuberculosis and pathogenesis has renewed interest in this area of research. Previous experimental studies were limited to the analysis of individual genes or enzymes, whereas recent advances in computational systems biology and high-throughput experimental technologies now allows metabolism to be studied on a genome scale. In the present article, we discuss the progress being made in applying system-level approaches to study the metabolism of this important pathogen.
An understanding of the dynamics of the metabolic profile of a bacterial cell is sought from a dynamical systems analysis of kinetic models. This modelling formalism relies on a deterministic mathematical description of enzyme kinetics and their metabolite regulation. However, it is severely impeded by the lack of available kinetic information, limiting the size of the system that can be modelled. Furthermore, the subsystem of the metabolic network whose dynamics can be modelled is faced with three problems: how to parameterize the model with mostly incomplete steady state data, how to close what is now an inherently open system, and how to account for the impact on growth. In this study we address these challenges of kinetic modelling by capitalizing on multi-‘omics’ steady state data and a genome-scale metabolic network model. We use these to generate parameters that integrate knowledge embedded in the genome-scale metabolic network model, into the most comprehensive kinetic model of the central carbon metabolism of E. coli realized to date. As an application, we performed a dynamical systems analysis of the resulting enriched model. This revealed bistability of the central carbon metabolism and thus its potential to express two distinct metabolic states. Furthermore, since our model-informing technique ensures both stable states are constrained by the same thermodynamically feasible steady state growth rate, the ensuing bistability represents a temporal coexistence of the two states, and by extension, reveals the emergence of a phenotypically heterogeneous population.
The structure and distribution of 1S990, a new Mycobacterium tuberculosis DNA sequence with homology to characterized insertion sequences (ISs), were investigated. IS990 was related to IS elements of the IS3 family and was present as a single copy in all 21 investigated M. tuberculosis strains, two Mycobacterium bovis strains and two M. bovis BCG strains. The sequence appears to be specific for the M. tuberculosis complex. The element carries two frameshift mutations and appears to be defective.
Background/Aims: Recent studies in primary biliary cirrhosis have reported the detection of serum antibodies against Mycobacterium gordonae and of myco-bacterial DNA in liver sections. The aim of this study was to investigate whether mycobacterial DNA is present in liver biopsy material in primary biliary cirrhosis. Methods: Archival liver biopsy specimens from 11 patients with primary biliary cirrhosis (10 female, mean age 52 years) and 11 patients with autoimmune hepatitis (10 female, mean age 53 years) were identified. Positive control tissue comprised five archival lymph node specimens from patients with tuberculous lymph-adenopathy, three of which had stained positive on ZN staining, and also a liver biopsy specimen from a patient with tuberculous hepatitis (ZN positive). Fixed sections were deparaffinised and DNA was extracted by mechanical disruption with glass beads. DNA was purified by use of diatoms and lysis in guanidinium thiocyanate in a technique previously validated for archival DNA. Primers were directed to amplify a partial 16S ribosomal RNA gene yielding the species-specific character for mycobacteria, and also to amplify the constitutively-expressed human gene GAPDH. Results: The polymerase chain reaction was shown to be capable of detecting 1 fg of M. gordonae DNA in ‘spiked’ samples, equivalent to 1–5 bacterial cells. No mycobacterial DNA was detected in liver biopsy samples from either the primary biliary cirrhosis or autoimmune hepatitis groups. Of the tuberculous control sections, mycobacterial DNA was detected in four of five lymph nodes and the liver biopsy specimen. GAPDH amplification was detected in all tested samples from liver disease and tuberculous control samples. Conclusion: These data do not support a role for mycobacteria in the aetiology of primary biliary cirrhosis.
Rapid, reliable, sensitive, portable, and accurate diagnostics are required to control disease outbreaks such as COVID-19 that pose an immense burden on human health and the global economy. Here we developed a loop-mediated isothermal amplification (LAMP)-based electrochemical test for the detection of SARS-CoV-2 that causes COVID-19. The test is based on the oxidation-reduction reaction between pyrophosphates (generated from positive LAMP reaction) and molybdate that is detected by cyclic voltammetry using inexpensive and disposable carbon screen printed electrodes. Our test showed higher sensitivity (detecting as low as 5.29 RNA copies/μL) compared to the conventional fluorescent reverse transcriptase (RT)-LAMP. We validated our tests using human serum and saliva spiked with SARS-CoV-2 RNA and clinical (saliva and nasal-pharyngeal) swab samples demonstrating 100% specificity and 93.33% sensitivity. Our assay provides a rapid, specific, and sensitive test with an electrochemical readout in less than 45 min that could be adapted for point-of-care settings.
Single walled carbon nanotubes (SWCNT) have attracted a great interest due their extraordinary properties that envisage their use for a wide range of applications [reference physical properties]. However, these properties are controlled by the chirality of the SWC-NTs. Unfortunately, the growth processes available to-date produce SWCNTs with different chiralities. Also, the SWCNTs are produced together with a relatively high quantities of impurities such as amorphous carbon and metallic catalyst particles. Indeed, the purification and manipulation remains problematic, hindering some of the possible applications of these materials. In this paper, the purification of SWCNTs with biological polymers is presented. The results shown that DNA and UNA effectively purify SWCNT from the "soot" obtained during the growth process. The results show how effectively total genomic UNA (tgRNA) purifies SWCNT. Atomic force microscopy (AFM) studies reveal how nucleic acids wrap around SWCNTs forming RNA-CNT composites. Moreover, when a RNA-CNT solution is dried on a hydrophilic surface, SWCNTs are found lying or embedded on a self assembled two dimensional UNA network. Using tgRNA is not only a cheap and effective method of solubilising and purifying CNTs but offers a first step towards the self-assembly of CNTs from solution. Furthermore, tgRNA networks could be a convenient method of electrically linking individual RNA functionalised CNTs over a surface which could prove useful for RNA or DNA biosensors.
Examination of Neisseria meningitidis strains associated with endemic meningococcal disease demonstrated differences in the number of copies of a repetitive sequence. Characterization of a copy of this repetitive sequence present in B15 strains has revealed the presence of a novel insertion sequence (IS1106) located within a complex repetitive region downstream of the gene for the major surface antigen (porA). IS1106 has a length of 1137 bp and is flanked by 36bp inverted repeats. Two open reading frames (ORF1 and ORF2) are present in opposite strands in codon-codon register with ORF2 entirely located within ORF1. The predicted protein from ORF1 demonstrates homology with the 5A protein of IS5 (Kroger and Hobom, 1982). Strains from two independent outbreaks of B15 meningococcal disease in the UK were found to contain the same genomic deletion removing a copy of IS 1 106 downstream of the porA gene.
Most healthy adults are protected from meningococcal disease by the presence of naturally acquired anti-meningococcal antibodies; however, the identity of the target antigens of this protective immunity remains unclear, particularly for protection against serogroup B disease. To identify the protein targets of natural protective immunity we developed an immunoprecipitation and proteomics approach to define the immunoproteome of the meningococcus. Sera from 10 healthy individuals showing serum bactericidal activity against both a meningococcal C strain (L91543) and the B strain MC58, together with commercially available pooled human sera were used as probe anti-sera. Immunoprecipitation was performed with each serum sample and live cells from both meningococcal strains. Immunoprecipitated proteins were identified by mass spectrometry. Analysis of the immunoproteome from each serum demonstrated both pan-reactive antigens that were recognised by most sera as well as subject-specific antigens. Most antigens were found in both meningococcal strains but a few were strain-specific. Many of the immunoprecipitated proteins have previously been characterised as surface antigens including adhesins and proteases, several of which have been recognised as vaccine candidate antigens e.g. fHBP, NadA and NHBA. The data clearly demonstrates the presence of meningococcal antibodies in healthy individuals with no history of meningococcal infection and a wide diversity of immune responses. The identification of the immuno-reactive proteins of the meningococcus provides a basis for understanding the role of each antigen in the natural immunity associated with carriage and may help to design vaccination strategies.
Strains of the Mycobacterium avium intracellulare complex (MAIC) have become important colonisers of patients with acquired immunodeficiency syndrome (AIDS). Restriction fragment length polymorphisms were used to study the DNA from 88 MAIC isolates, including 51 derived from 47 AIDS patients. MAIC isolates from 33 of 45 AIDS patients were identical at the molecular level and distinct from the mycobacteria isolated from the stools of healthy subjects. The study also showed that serotyping correlates poorly with the genetic identity of these organisms. Mycobacterium paratuberculosis, which has been implicated in Crohn's disease, was not identified in any of the cultures studied.
Evolution within a rugged fitness landscape is limited by the tendency for organisms to become trapped on local optima resulting in evolutionary stasis. It is presently unclear how founder populations escape from an adaptive peak to found a new species. Insertion sequences, transposons and other mobile DNA elements are found in all species of eukaryotes, bacteria and archaebacteria, where they have been sought and are usually considered to be genomic parasites or selfish genes, However, many transposons and other mobile repetitive DNA are remarkably species or phyla-specific, indicating that infection with transposable elements coincides with speciation events and is involved in promoting evolutionary change. We propose here a model in which transposable elements are involved in speciation events by their ability to produce irreversible deleterious mutations that promote escape from evolutionary stasis. We have constructed a genetic algorithm designed to model both spontaneous and transposon-mediated mutations in populations of asexual digital organisms. We use this model to investigate the effect of transposon-mediated mutations on the rate of evolution of digital organisms as they compete for resources within an artificial adaptive landscape. In the absence of transposon mutations the seed organisms quickly evolve to occupy the nearest adaptive peak but thereafter evolutionary stasis ensues and adjacent empty peaks are left unoccupied. In the presence of transposon mutations, evolution is again dominated by stasis but is punctuated by bursts of rapid evolution in which consecutive unoccupied adaptive peaks are filled with organisms derived from single transposition events. Rapid evolutionary events leading to founding of new biological species, may be similarly initiated by irreversible deleterious mutations induced by transposition.
The Sm14 antigen of Schistosoma mansoni was cloned and expressed in Mycobacterium bovis BCG as a fusion with the Mycobacterium fortuitum P-lactamase protein under the control of its promoter, pBlaF*; the protein was localized in the bacterial cell wall. The rBCG-Sm14 strain was shown to be relatively stable in cultured murine and bovine monocytes in terms of infectivity, bacterial persistence, and plasmid stability. The immunization of mice with rBCG-Sm14 showed no induction of anti-Sm14 antibodies; however, splenocytes of immunized mice released increased levels of gamma interferon upon stimulation with recombinant Sm14 (rSm14), indicating an induction of a Th1-predominant cellular response against Sm14. Mice immunized with one or two doses of rBCG-Sm14 and challenged with live S. mansoni cercaria showed a 48% reduction in worm burden, which was comparable to that obtained by immunization with three doses of rSm14 purified from Escherichia coli. The data presented here further enhance the status of Sm14 as a promising candidate antigen for the control of schistosomiasis and indicate that a one-dose regimen of rBCG-Sm14 could be considered a convenient means to overcome many of the practical problems associated with the successful implementation of a multiple-dose vaccine schedule in developing countries.
Cell growth experiments with a microfluidic device produce large scale time-lapse image data, which contain important information on cell growth and patterns in their genealogy. To extract such information, we propose a scheme to segment and track bacterial cells automatically. In contrast to most published approaches, which often split segmentation and tracking into two independent procedures, we focus on designing an algorithm that describes cell properties evolving between consecutive frames by feeding segmentation and tracking results from one frame to the next one. The cell boundaries are extracted by minimising the Distance Regularised Level Set Evolution model. Each individual cell was identified and tracked by identifying cell septum and membrane as well as developing a trajectory energy minimisation function along time-lapse series. Experiments show that by applying this scheme, cell growth and division can be measured automatically. The results show the efficiency of the approach when testing on different datasets while comparing with other existing algorithms. The proposed approach demonstrates great potential for large scale bacterial cell growth analysis.
The principle that mutations occur randomly with respect to the direction of evolutionary change has been challenged by the phenomenon of adaptive mutations. There is currently no entirely satisfactory theory to account for how a cell can selectively mutate certain genes in response to environmental signals. However, spontaneous mutations are initiated by quantum events such as the shift of a single proton (hydrogen atom) from one site to an adjacent one. We consider here the wave function describing the quantum state of the genome as being in a coherent linear superposition of states describing both the shifted and unshifted protons. Quantum coherence will be destroyed by the process of decoherence in which the quantum state of the genome becomes correlated (entangled) with its surroundings. Using a very simple model we estimate the decoherence times for protons within DNA and demonstrate that quantum coherence may be maintained for biological time-scales. Interaction of the coherent genome wave function with environments containing utilisable substrate will induce rapid decoherence and thereby destroy the superposition of mutant and non-mutant states. We show that this accelerated rate of decoherence may significantly increase the rate of production of the mutated state.
An in-depth study of a novel functionalization of carbon nanotubes for their application as protein and DNA carriers is presented. First, the optimum conditions for the dispersion of singlewalled carbon nanotubes (SWCNTs) with amphiphilic polypeptides were obtained, and the SWCNT–polypeptide complexes were characterized by different techniques (UV–Vis-NIR, CD, and AFM). Based on the properties of the SWCNT–polypeptide complexes, a model that characterizes the adsorption of natural proteins onto SWCNT was described for the first time. This model predicts the adsorption of natural proteins on SWCNTs based on the protein structure and composition, and therefore, allows the design of methods for the preparation of SWCNT–protein complexes. Besides, the use of cationic-designed amphiphilic polypeptides to disperse SWCNTs is applied for subsequent and efficient binding of DNA to carbon nanotubes by a bilayer approach. Therefore, in this article, we develop procedures for the use of SWCNTs as protein and DNA carriers. The systems were delivered into cells showing that the efficiency of delivery is affected by the charge of the complexes, which has important implications in the use of SWCNT as platforms for protein and DNA binding and subsequent use as delivery systems.
pUS933, a bifunctional Mycobacterium-Escherichia coli translational fusion vector containing an amino-terminally truncated E. coli lacZ reporter gene, was constructed. Derivatives of pUS933, containing the promoter, RBS and start codon of the Mycobacterium bovis BCG hsp60 gene, the Mycobacterium leprae 28 kDa gene and the M. leprae 18 kDa gene were constructed and introduced into E. coli, Mycobacterium smegmatis and M. bovis BCG. beta-Galactosidase activity was measured for mycobacteria grown in liquid culture. Primer-extension analysis was used to determine the transcriptional start point for the 18 kDa promoter in M. smegmatis. Murine macrophages were infected with recombinant BCG containing the pUS933 derivatives and expression levels were examined, by fluorescence microscopy and fluorometry, during intracellular growth of BCG. Both the BCG hsp60 gene promoter and the M. leprae 28 kDa gene promoter gave high levels of beta-galactosidase expression in all situations examined. In contrast, the M. leprae 18 kDa promoter fragment gave very low levels of expression in M. smegmatis and BCG grown in liquid culture, but in BCG growing within macrophages it was induced to levels almost as high as the other promoters. This indicated that the 18 kDa gene is specifically activated during intracellular growth and may therefore be involved in survival of M. leprae within macrophages. This pattern of regulation may be useful for controlling expression of foreign genes in recombinant BCG strains.
An artificial mycobacterial transposon was constructed by placing two copies of the insertion sequence IS900 flanking a kanamycin resistance gene into a non-(mycobacterial) replicating vector. Constructs were introduced into mycobacteria by electroporation and transposition events conferring kanamycin resistance were selected. Integration of IS900 into several genomic sites was analysed by Southern blotting and shown to involve both simple insertions and cointegrate formation, suggesting that IS900 can transpose by a replicative mechanism. Kanamycin resistance of IS900-integrated transformants was shown to be stable in the absence of selection.
Despite decades of research many aspects of the biology of Mycobacterium tuberculosis remain unclear and this is reflected in the antiquated tools available to treat and prevent tuberculosis and consequently this disease remains a serious public health problem. Important discoveries linking M. tuberculosis’s metabolism and pathogenesis have renewed interest in this area of research. Previous experimental studies were limited to the analysis of individual genes or enzymes whereas recent advances in computational systems biology and high throughput experimental technologies now allow metabolism to be studied on a genome scale. Here we discuss the progress being made in applying system level approaches to studying the metabolism of this important pathogen. The information from these studies will fundamentally change our approach to tuberculosis research and lead to new targets for therapeutic drugs and vaccines.
Mycobacterium vaccae represents an alternative mycobacterial cloning host that has been largely overlooked to date. The main reason for this may be the reported non-transformability of this species, specifically the so-called Stanford strain (NCTC 11659), with expression vectors that use kanamycin resistance as a selection method. However, this strain can be transformed using hygromycin resistance as an alternative selectable phenotype. The present study has shown that in contrast to previous reports, M. vaccae (ATCC 15483) is capable of being transformed with a range of vectors encoding kanamycin resistance as the selectable marker. Thereafter, the expression of the lacZ reporter gene in M. vaccae, Mycobacterium bovis BCG and Mycobacterium smegmatis mc(2)155 was evaluated using a range of characterized mycobacterial promoter sequences (hsp60, hsp70, PAN, 18kDa and 16S rRNA) cloned in the same promoter probe vector. In general, the promoters showed similar levels of activity in the three species, demonstrating that existing expression systems can readily be employed with M. vaccae (ATCC 15483). This was further confirmed by the observation that M. vaccae was capable of stable, in vitro expression of recombinant S1 subunit of pertussis toxin at levels equivalent to those obtained with BCG and M. smegmatis. Analysis of structural and functional stability of a range of vectors demonstrated that the incidence of instability noted for M. vaccae was lower than that recorded for M. smegmatis. Taken together, the results indicate that M. vaccae is an additional cloning host which may prove useful for specific aspects of mycobacterial biology and provide increased flexibility to the field of recombinant protein technology for mycobacteria.
Leprosy, caused by Mycobacterium leprae, has plagued humanity for thousands of years and continues to cause morbidity, disability and stigmatization in two to three million people today. Although effective treatment is available, the disease incidence has remained approximately constant for decades so new approaches, such as vaccine or new drugs, are urgently needed for control. Research is however hampered by the pathogen’s obligate intracellular lifestyle and the fact that it has never been grown in vitro. Consequently, despite the availability of its complete genome sequence, fundamental questions regarding the biology of the pathogen, such as its metabolism, remain largely unexplored. In order to explore the metabolism of the leprosy bacillus with a long-term aim of developing a medium to grow the pathogen in vitro, we reconstructed an in silico genome scale metabolic model of the bacillus, GSMN-ML. The model was used to explore the growth and biomass production capabilities of the pathogen with a range of nutrient sources, such as amino acids, glucose, glycerol and metabolic intermediates. We also used the model to analyze RNA-seq data from M. leprae grown in mouse foot pads, and performed Differential Producibility Analysis to identify metabolic pathways that appear to be active during intracellular growth of the pathogen, which included pathways for central carbon metabolism, co-factor, lipids, amino acids, nucleotides and cell wall synthesis. The GSMN-ML model is thereby a useful in silico tool that can be used to explore the metabolism of the leprosy bacillus, analyze functional genomic experimental data, generate predictions of nutrients required for growth of the bacillus in vitro and identify novel drug targets.
New approaches are needed to control leprosy, but understanding of the biology of the causative agent Mycobacterium leprae remains rudimentary, principally because the pathogen cannot be grown in axenic culture. Here, we applied 13C isotopomer analysis to measure carbon metabolism of M. leprae in its primary host cell, the Schwann cell. We compared the results of this analysis with those of a related pathogen, Mycobacterium tuberculosis, growing in its primary host cell, the macrophage. Using 13C isotopomer analysis with glucose as the tracer, we show that whereas M. tuberculosis imports most of its amino acids directly from the host macrophage, M. leprae utilizes host glucose pools as the carbon source to biosynthesize the majority of its amino acids. Our analysis highlights the anaplerotic enzyme phosphoenolpyruvate carboxylase required for this intracellular diet of M. leprae, identifying this enzyme as a potential antileprosy drug target.
Summary We have constructed a mycobacterial integrative vector by placing two copies of the insertion sequence IS900 flanking a kanamycin‐resistance gene into a 'suicide’vector unable to replicate in mycobacteria. The Mycobacterium leprae gene encoding the M. leprae 18kDa protein was cloned between the two copies of IS900 to provide expression signals. Constructs were introduced into Mycobacterium species smegmatis, vaccae and bovis BCG by electroporation and selection for kanamycin resistance. The expression of the 18 kDa gene was analysed by Western blotting. Integration of the vector into the M. smegmatis chromosome was analysed by Southern blotting. One to five copies of the vector were detected in each transformant. The SIV gag p27 gene and the foot‐and‐mouth disease virus VP1 140‐160 epitope were successfully cloned into the 18kDa gene and expression in M. smegmatis was obtained.
Expression vectors containing rabies virus nucleoprotein B-cell and T-cell epitopes in Mycobacterium bovis BCG were constructed. The epitopes were subcloned into the M. leprae 18-kDa gene to ensure correct presentation to the host immune system. Expression of the 18-kDa::B+T epitope fusion protein was driven by either the hsp60 promoter, which is constitutively activated at a high level in M. bovis BCG, or the 18-kDa promoter, which is strongly induced in vivo. Mice were immunised intra-peritoneally with the recombinant BCG cultures and compared to a control group vaccinated with the commercial rabies vaccine Rai-SAD. Both of the expression vectors elicited a higher antibody titre than that of the rabies vaccine, with the highest response shown by M. bovis BCG (pUP203), expression controlled by the 18-kDa promoter. Immunisation with M. bovis BCG (pUP202), expression controlled by the hsp60 promoter, resulted in a continuously increasing antibody titre up to 60 days post immunisation. The mice antibodies were also capable of recognising the whole rabies virus and not only the synthetic peptide epitopes.
To investigate the epidemiology of meningococcal disease, a specific DNA probe pUS210 (carrying insert DNA which is repeated in the meningococcal genome) was isolated. The ability of this probe to hydridise with multiple polymorphic fragments in Southern blots was exploited to examine genetic relations within strains. Two geographically distinct foci of prolonged meningococcal disease (Gloucester and Plymouth, UK) are due to a clonal population of virulent strains that are distinct from those found elsewhere in the UK.
Neisseria meningitidis is a global cause of meningitis and septicemia. Immunity to N. meningitidis involves both innate and specific mechanisms with killing by serum bactericidal activity and phagocytic cells. C-reactive protein (CRP) is an acute-phase serum protein that has been shown to help protect the host from several bacterial pathogens, which it recognizes by binding to phosphorylcholine (PC) on their surfaces. Pathogenic Neisseria species can exhibit phase-variable PC modification on type 1 and 2 pili. We have shown that CRP can bind to piliated meningococci in a classical calcium-dependent manner. The binding of CRP to the meningococcus was concentration dependent, of low affinity, and specific for PC. CRP appears to act as an opsonin for N. meningitidis, as CRP-opsonized bacteria showed increased uptake by human macrophages and neutrophils. Further investigation into the downstream effects of CRP-bound N. meningitidis may lead us to a better understanding of meningococcal infection and help direct more effective therapeutic interventions.
Whereas intracellular carbon metabolism has emerged as an attractive drug target, the carbon sources of intracellularly replicating pathogens, such as the tuberculosis bacillus Mycobacterium tuberculosis, which causes long-term infections in one-third of the world's population, remain mostly unknown. We used a systems-based approach-(13)C-flux spectral analysis (FSA) complemented with manual analysis-to measure the metabolic interaction between M. tuberculosis and its macrophage host cell. (13)C-FSA analysis of experimental data showed that M. tuberculosis obtains a mixture of amino acids, C1 and C2 substrates from its host cell. We experimentally confirmed that the C1 substrate was derived from CO2. (13)C labeling experiments performed on a phosphoenolpyruvate carboxykinase mutant revealed that intracellular M. tuberculosis has access to glycolytic C3 substrates. These findings provide constraints for developing novel chemotherapeutics.
A mycobacterial aetiology for Crohn's disease (CD) has been suggested. Slow growing mycobacteria indistinguishable from M paratuberculosis, the causative agent of enteritis in ruminants (Johne's disease) have been isolated from CD tissues. We have used cloned genomic DNA probes derived from a CD isolated mycobacteria strain Ben, to investigate the presence of mycobacterial DNA sequences in CD tissues. DNA was extracted from total tissue from 17 CD and four control gut specimens. DNA was digested with restriction endonucleases, electrophoresed and transferred to nylon membranes by Southern blotting and hybridised to radiolabelled DNA probes. No mycobacterial DNA was detected in any tissue sample studied. Reconstitution experiments with known numbers of in vitro cultured mycobacteria showed sensitive detection of mycobacterial DNA. DNA extracted from mouse liver, infected with M lepraemurium revealed a strong hybridisation signal and showed the applicability of the experimental approach to the detection of mycobacterial DNA in naturally infected tissues. The results do not provide evidence for the involvement of mycobacteria in the pathogenesis of CD but do not exclude the possibility of low levels of infection in subsets of intestinal cells with spheroplast or cell wall deficient forms of mycobacteria.
The conscious electromagnetic information (cemi) field theory proposes that the seat of consciousness is the brain’s electromagnetic (EM) field that integrates information from trillions of firing neurons. What we call free will is its output. The cemi theory also proposes that the brain has two streams. Most actions are initiated by the first non-conscious stream that is composed of neurons that are insulated from EM field influences. These non-conscious involuntary actions are thereby invisible to our EM field-located thoughts. The theory also proposes that voluntary actions are driven by neurons that receive EM field inputs and are thereby visible to our EM field-located thoughts. I review the extensive evidence for EM field/ephaptic coupling between neurons and the increasing evidence that EM fields in the brain are a cause of behaviour. I conclude by arguing that though this EM field-driven will is not free, in the sense of being acausal, it nevertheless corresponds to the very real experience of our conscious mind being in control of our voluntary actions. Will is not an illusion. It is our experience of control by our EM field-located mind. It is an immaterial, yet physical, will.
We have designed a drug delivery system for the anti-cancer drugs doxorubicin and mitoxantrone based on carbon nanotubes, which is stable under biological conditions, allows for sustained release, and promotes selectivity through an active targeting scheme. Carbon nanotubes are particularly promising for this area of application due to their high surface area, allowing for high drug loading, and their unique interaction with cellular membranes. We have taken a systematic approach to PEG conjugation in order to create a formulation of stable and therapeutically effective CNTs. The presented drug delivery system may be a means of improving cancer treatment modalities by reducing drug-related side effects.
Quantum biology is usually considered to be a new discipline, arising from recent research that suggests that biological phenomena such as photosynthesis, enzyme catalysis, avian navigation or olfaction may not only operate within the bounds of classical physics but also make use of a number of the non-trivial features of quantum mechanics, such as coherence, tunnelling and, perhaps, entanglement. However, although the most significant findings have emerged in the past two decades, the roots of quantum biology go much deeper—to the quantum pioneers of the early twentieth century. We will argue that some of the insights provided by these pioneering physicists remain relevant to our understanding of quantum biology today.
Polyethyleneimine-coated double-walled carbon nanotubes (DWCNTs) were used for dual gene and drug delivery, after loading the DWCNTs with the drug chloroquine, a lysosomotropic compound that is able to promote escape from the lysosomal compartment. Different forms of functionalization of the DWCNTs were examined in order to optimize this system. They included the testing of different treatments on DWCNTs to optimize the loading and delivery of chloroquine and the selection of a cationic polymer for coating the DWCNTs for optimum DNA binding and delivery. An acid oxidation treatment of DWCNTs was selected for optimum chloroquine loading together with polyethyleneimine as optimum cationic coating agent for plasmid DNA binding. Optimization of the conditions for choroquine-enhanced gene delivery were developed using luciferase expression as a model system. We have demonstrated that chloroquine-loading increases the ability of polyethyleneimine-coated DWCNTs to deliver functional nucleic acid to human cells. Cell viability tests have shown no cytotoxicity of the functionalized DWCNTs at the concentrations needed for optimum gene delivery. These results support the potential applications of this methodology in gene therapy.
The rational use of IS6110 fingerprinting for studies of the molecular epidemiology and evolution of Mycobacterium tuberculosis requires understanding of the dynamics of transposition. In laboratory model systems, it has been shown that transposition is context-sensitive, i.e. it is influenced by the nature of the site in which the insertion sequence is presented. Stimulation of transposition by activation of an adjacent promoter supports the hypothesis that transposition occurs more readily from transcriptionally active locations. In addition, it has been shown that transposition can be enhanced by the expression of the transposase in trans. These findings imply that the frequency of transposition will vary substantially between different strains of M. tuberculosis, and furthermore that a hitherto stable strain may develop more rapid variation due to transposition into an active site. The use of IS6110 fingerprinting for the analysis of longer-range relationships between M. tuberculosis isolates therefore needs to be interpreted with care.
In earlier papers I described the conscious electromagnetic information (CEMI) field theory, which claimed that the substrate of consciousness is the brain's electromagnetic (EM) field. I here further explore this theory by examining the properties and dynamics of the information underlying meaning in consciousness. I argue that meaning suffers from a binding problem, analogous to the binding problem described for visual perception, and describe how the gestalt (holistic) properties of meaning give rise to this binding problem. To clarify the role of information in conscious meaning, I differentiate between extrinsic information that is symbolic and arbitrary, and intrinsic information, which preserves structural aspects of the represented object and thereby maintains some gestalt properties of the represented object. I contrast the requirement for a decoding process to extract meaning from extrinsic information, whereas meaning is intrinsic to the structure of the gestalt intrinsic information and does not require decoding. I thereby argue that to avoid the necessity of a decoding homunculus, conscious meaning must be encoded intrinsically -- as gestalt information -- in the brain. Moreover, I identify fields as the only plausible substrate for encoding gestalt intrinsic information and argue that the binding problem of meaning can only be solved by grounding meaning in this field-based gestalt information. I examine possible substrates for gestalt information in the brain and conclude that the only plausible substrate is the CEMI field.
: It's been 20 years since the first report of a recombinant vaccine that protected against leptospirosis. Since then, numerous recombinant vaccines have been evaluated; however, no recombinant vaccine candidate has advanced to clinical trials. With the ever-increasing burden of leptospirosis, there is an urgent need for a universal vaccine against leptospirosis. : This review covers the most promising vaccine candidates that induced significant, reproducible, protection and how advances in the field of bioinformatics has led to the discovery of hundreds of novel protein targets. The authors also discuss the most recent findings regarding the innate immune response and host-pathogen interactions and their impact on the discovery of novel vaccine candidates. In addition, the authors have identified what they believe are the most challenging problems for the discovery and development of a universal vaccine and their potential solutions. : A universal vaccine for leptospirosis will likely only be achieved using a recombinant vaccine as the bacterins are of limited use due to the lack of a cross-protective immune response. Although there are hundreds of novel targets, due to the lack of immune correlates and the need for more research into the basic microbiology of spp., a universal vaccine is 10-15 years away.
The principles of non-covalent functionalization of carbon nanotubes will be described. The abilities of these biomolecules to solubilize carbon nanotubes and bind DNA will be also compared. Approaches for using functionalized carbon nanotubes to deliver genes to target cells and associated problems will be described. Evidence pertaining to the mechanism of entry of nucleic acid-loaded carbon nanotubes into mammalian cells will be also presented.
An enzymic method for the extraction of high molecular weight DNA from a wide range of diverse bacteria, including mycobacteria, is described. The method is simple, rapid, requires few manipulations and is suitable for extracting quantitative yields of DNA from as few as ≈3×10 7 bacterial cells. The DNA produced is of good quality and is suitable for molecular biological manipulations.
To detect Mycobacterium tuberculosis in clinical samples, we used the M. tuberculosis–complex specific insertion sequence IS 990 as the target in a simple DIG-PCR ELISA assay, as this element is present as a single copy in all strains of M. tuberculosis we have examined to date. The IS 990 test was compared with a similar PCR that utilizes IS 6110 as target. For detection of PCR product, digoxigenin-11-dUTP (DIG-dUTP) was incorporated into the product. After amplification, the PCR product was hybridized with biotinylated capture probe, which was complementary to the inner part of the amplicon. The hybrid was captured onto streptavidin-coated microtiter plate and DIG-labeled PCR product was detected using a peroxidase-conjugated antibody to DIG. We evaluated DIG-PCR ELISA for the detection ofM. tuberculosis DNA in 265 respiratory and non-respiratory specimens taken from patients with known and suspected tuberculosis disease or from controls. The sensitivity and specificity of both IS 990 -based test and IS 6110 -based test was 96.5% and 95.3% respectively, comparable to the sensitivity and specificity of the IS 6110 -based test. The results demonstrate that the IS 990 PCR ELISA test is a rapid and sensitive tool for the detection and identification of M. tuberculosis in clinical samples, and may have advantages to the more widely used IS 6110 -based tests, particularly in areas where IS 6110 -negative strains are found.
Constraint-based approaches facilitate the prediction of cellular metabolic capabilities, based, in turn on predictions of the repertoire of enzymes encoded in the genome. Recently, genome annotations have been used to reconstruct genome scale metabolic reaction networks for numerous species, including Homo sapiens, which allow simulations that provide valuable insights into topics, including predictions of gene essentiality of pathogens, interpretation of genetic polymorphism in metabolic disease syndromes and suggestions for novel approaches to microbial metabolic engineering. These constraint-based simulations are being integrated with the functional genomics portals, an activity that requires efficient implementation of the constraint-based simulations in the web-based environment.
Introduction: Antibiotic persistence (subpopulation tolerance) occurs when a subpopulation of antibiotic sensitive cells survives prolonged exposure to a bactericidal concentration of an antibiotic, and is capable of regrowth once the antibiotic is removed. This phenomenon has been shown to contribute to prolonged treatment duration, infection recurrence, and accelerated development of genetic resistance. Currently, there are no biomarkers which would allow for segregation of these antibiotic-tolerant cells from the bulk population prior to antibiotic exposure, limiting research on this phenomenon to retrograde analyses. However, it has been previously shown that persisters often have a dysregulated intracellular redox homeostasis, warranting its investigation as a potential marker for antibiotic tolerance. Furthermore, it is currently unknown whether another antibiotic tolerant subpopulation - viable but non-culturable cells (VBNCs), are simply persisters with extreme lag phase, or are formed through separate pathways. VBNCs similarly to persisters remain viable following antibiotic exposure, however, are not capable of regrowth in standard conditions. Methods: In this article we employed an NADH:NAD+ biosensor (Peredox) to investigate NADH homeostasis of ciprofloxacin-tolerant E. coli cells on a single-cell level. [NADH:NAD+] was used as a proxy for measuring intracellular redox homeostasis and respiration rate. Results and Discussion: First, we demonstrated that ciprofloxacin exposure results in a high number of VBNCs, several orders of magnitude higher than persisters. However, we found no correlation in the frequencies of persister and VBNC subpopulations. Ciprofloxacin-tolerant cells (persisters & VBNCs) were actively undergoing respiration, although at a significantly lower rate on average when compared to the bulk population. We also noted significant heterogeneity on a single-cell level within the subpopulations, however were unable to segregate persisters from VBNCs based on these observations alone. Finally, we showed that in the highly-persistent strain of E. coli, E. coli HipQ, ciprofloxacin-tolerant cells have a significantly lower [NADH:NAD+] ratio than tolerant cells of its parental strain, providing further link between disturbed NADH homeostasis and antibiotic tolerance.
Biological effects of electromagnetic fields (EMFs) have previously been identified for cellular proliferation and changes in expression and conduction of diverse types of ion channels. The major effect elicited by EMFs seems to be directed toward Ca2+ homeostasis. This is particularly remarkable since Ca2+ acts as a central modulator in various signaling pathways, including, but not limited to, cell differentiation and survival. Despite this, the mechanisms underlying this modulation have yet to be unraveled. Here, we assessed the effect of EMFs on intracellular [Ca2+], by exposing HEK 293 cells to both radio-frequency electromagnetic fields (RF-EMFs) and static magnetic fields (SMFs). We detected a constant and significant increase in [Ca2+] subsequent to exposure to both types of fields. Strikingly, the increase was nulled by administration of 10 mu M Thapsigargin, a blocker of sarco/endoplasmic reticulum Ca2+-ATPases (SERCAs), indicating the involvement of the endoplasmic reticulum (ER) in EMF-related modulation of Ca2+ homeostasis.
Many aspects of chemistry and biology are mediated by electromagnetic field (EMF) interactions. The central nervous system (CNS) is particularly sensitive to EMF stimuli. Studies have explored the direct effect of different EMFs on the electrical properties of neurons in the last two decades, particularly focusing on the role of voltage-gated ion channels (VGCs). This work aims to systematically review published evidence in the last two decades detailing the effects of EMFs on neuronal ion channels as per the PRISM guidelines. Following a predetermined exclusion and inclusion criteria, 22 papers were included after searches on three online databases. Changes in calcium homeostasis, attributable to the voltage-gated calcium channels, were found to be the most commonly reported result of EMF exposure. EMF effects on the neuronal landscape appear to be diverse and greatly dependent on parameters, such as the field's frequency, exposure time, and intrinsic properties of the irradiated tissue, such as the expression of VGCs. Here, we systematically clarify how neuronal ion channels are particularly affected and differentially modulated by EMFs at multiple levels, such as gating dynamics, ion conductance, concentration in the membrane, and gene and protein expression. Ion channels represent a major transducer for EMF-related effects on the CNS.
Carbon nanotubes (CNTs) exhibit unique size, shape and physical properties, which make them promising candidates for biomedical applications. In particular, carbon nanotubes have been intensively studied for conjugation with pre-existing therapeutic agents for more effective targeting, as a result of their ability to cross cell membranes. However, to utilise them effectively in biological systems it is extremely important to understand how they behave at the cellular level and their distribution in vivo. Additionally, in order to consider carbon nanotubes as candidate delivery systems of therapeutic agents it is important to ascertain their fate in vivo, but also take into account many factors, such as solubility, stability and clearance. Issues surrounding their short term and long term safety are currently the subject of toxicology testing. Herein, we propose to summarize the main findings on the uptake, trafficking and biodistribution of carbon nanotubes, with special focus on functionalized carbon nanotubes for delivery of therapeutic agents.
We have used DNA and protein polymorphisms for the third complement component (C3) to assess the potential of DNA markers in the diagnosis and study of familial hypercholesterolaemia (FH), and to confirm the reported linkage between FH and C3. The inheritance of FH and the C3 gene has been studied in 10 families by combining information from both the protein and DNA polymorphisms. Our results confirm that the C3 gene is loosely linked to the gene causing FH (lod score maximum of 2.0) at a recombination distance of 0.15. When these results are combined with previously published data the overall lod score maximum is 4.75 at a recombination distance of 0.2, meaning that the two genes will be inherited together in only about 80% of children. These results confirm that the gene that causes familial hypercholesterolaemia is linked to C3 and is therefore on chromosome 19, but C3 is not close enough to be used as a diagnostic marker.
Objectives: To use genetic fingerprinting to investigate the epidemiology of tuberculosis (FB) caused by Mycobacterium tuberculosis in Poland, a country with a relatively high incidence of tuberculosis, to improve TB control. Design: One hundred M. tuberculosis isolates from 98 patients in the Institute of Tuberculosis and Lung Diseases in Warsaw from 1993 to 1995 and 85 isolates obtained from 50 patients in the Hospital of Lung Diseases in Lodz in 1996 were subjected to DNA restriction fragment length polymorphism (RFLP) analysis, using the insertion sequence IS6110 as a probe. Results: IS 6110-associated banding patterns of the M. tuberculosis isolates originating from different localities varied considerably, but isolates from Lodz had a higher degree of similarity. Strains with identical RFLP types were identified in patients of the same family or patients living in the same area, indicating active transmission. Of strains isolated in Warsaw, 45% were resistant to at least one drug, and 35% were resistant to two or more drugs and were classified as multidrugresistant ( MDR). Some drug-resistant isolates were found to have identical banding patterns and originated from epidemiologically linked cases. Conclusions: Active transmission of TB, including MDR TB, is occurring in Poland. Active measures must be taken to prevent the spread of drug-resistant TB in Poland and potentially, the rest of Europe.
Klebsiella pneumoniae is an important pathogenic bacterium commonly associated with human healthcare and community-acquired infections. In recent years, K. pneumoniae has become a significant threat to global public and veterinary health, because of its high rates of antimicrobial resistance (AMR). Early diagnosis of K. pneumoniae infection and detection of any associated AMR would help to accelerate directed therapy and reduce the risk of the emergence of multidrug-resistant isolates. In this study, we identified three target genes (yhaI, epsL, and xcpW) common to K. pneumoniae isolates from both China and Europe and designed loop-mediated isothermal amplification (LAMP) assays for the detection of K. pneumoniae in clinical samples. We also designed LAMP assays for the detection of five AMR genes commonly associated with K. pneumoniae. The LAMP assays were validated on a total of 319 type reference strains and clinical isolates of diverse genetic backgrounds, in addition to 40 clinical human sputum samples, and were shown to be reliable, highly specific, and sensitive. For the K. pneumoniae-specific LAMP assay, the calculated sensitivity, specificity, and positive and negative predictive values (comparison with culture and matrix-assisted laser desorption/ionization-time of flight mass spectrometry) were all 100% on clinical isolates and, respectively, of 100%, 91%, and 90%, and 100% when tested on clinical sputum samples, while being significantly faster than the reference methods. For the bla(KPC) and other carbapenemases' LAMP assays, the concordance between the LAMP results and the references methods (susceptibility tests) was 100%, on both pure cultures (n = 125) and clinical samples (n = 18). In conclusion, we developed highly sensitive and specific LAMP assays for the clinical identification of K. pneumoniae and detection of carbapenem resistance.
Brian D. Robertson and Brendan W. Wren
Efforts to develop carbon nanotubes (CNTs) as nano-vehicles for precise and controlled drug and gene delivery, as well as markers for in vivo biomedical imaging, are currently hampered by uncertainties with regard to their cellular uptake, their fate in the body, and their safety. All of these processes are likely to be affected by the purity of CNT preparation, as well as the size and concentration of CNTs used, parameters that are often poorly controlled in biological experiments. It is demonstrated herein that under the experimental conditions of standard transfection methods, DWNTs are taken up by cultured cells but are then released after 24 h with no discernable stress response. The results support the potential therapeutic use of CNTs in many biomedical settings, such as cancer therapy.
Neisseria meningitidis is the etiologic agent of meningococcal meningitis and sepsis. Initial colonization of meningococci in the upper respiratory tract epithelium is crucial for disease development. The colonization occurs in several steps and expression of type IV pili (Tfp) is essential for both attachment and microcolony formation of encapsulated bacteria. Previously, we have shown that host-derived lactate induces synchronized dispersal of meningococcal microcolonies. In this study, we demonstrated that lactate-induced dispersal is dependent on bacterial concentration but not on the quorum-sensing system autoinducer-2 or the two-component systems NarP/NarQ, PilR/PilS, NtrY/NtrX, and MisR/MisS. Further, there were no changes in expression of genes related to assembly, elongation, retraction, and modification of Tfp throughout the time course of lactate induction. By using pill and pptB mutants, however, we found that lactate-induced dispersal was dependent on PilT retraction but not on phosphoglycerol modification of Tfp even though the PptB activity was important for preventing reaggregation postdispersal. Furthermore, protein synthesis was required for lactate-induced dispersal. Finally, we found that at a lower temperature, lactate-induced dispersal was delayed and unsynchronized, and bacteria reformed microcolonies. We conclude that lactate-induced microcolony dispersal is dependent on bacterial concentration, PilT-dependent Tfp retraction, and protein synthesis and is influenced by environmental temperature.
Despite the introduction of conjugated polysaccharide vaccines for many of the Neisseria meningitidis serogroups, neisserial infections continue to cause septicaemia and meningitis across the world. This is in part due to the difficulties in developing a, cross-protective vaccine that is effective against all serogroups, including serogroup B meningococci. Although convalescent N. meningitidis patients develop a natural long-lasting cross-protective immunity, the antigens that mediate this response remain unknown. To help define the target of this protective immunity we identified the proteins recognized by IgG in sera from meningococcal patients by a combination of 2D protein gels, western blots and mass spectrometry. Although a number of outer membrane antigens were identified the majority of the antigens were cytoplasmic, with roles in cellular processes and metabolism. When recombinant proteins were expressed and used to raise sera in mice, none of the antigens elicited a positive SBA result, however flow cytometry did demonstrate that some, including the ribosomal protein, RplY were localised to the neisserial cell surface.
The use of carbon nanotubes as a gene delivery system has been extensively studied in recent years owing to its potential advantages over viral vectors. To achieve this goal, carbon nanotubes have to be functionalized to become compatible with aqueous media and to bind the genetic material. To establish the best conditions for plasmid DNA binding, we compare the dispersion properties of single-, double- and multi-walled carbon nanotubes (SWCNTs, DWCNTs and MWCNTs, respectively) functionalized with a variety of surfactants by non-covalent attachment. The DNA binding properties of the functionalized carbon nanotubes were studied and compared by electrophoresis. Furthermore, a bilayer functionalization method for DNA binding on SWCNTs was developed that utilized RNA-wrapping to solubilize the nanotubes and cationic polymers as a bridge between nanotubes and DNA.
Mycobacterium tuberculosis requires the enzyme isocitrate lyase (ICL) for growth and virulence in vivo. The demonstration that M. tuberculosis also requires ICL for survival during nutrient starvation and has a role during steady state growth in a glycerol limited chemostat indicates a function for this enzyme which extends beyond fat metabolism. As isocitrate lyase is a potential drug target elucidating the role of this enzyme is of importance; however, the role of isocitrate lyase has never been investigated at the level of in vivo fluxes. Here we show that deletion of one of the two icl genes impairs the replication of Mycobacterium bovis BCG at slow growth rate in a carbon limited chemostat. In order to further understand the role of isocitrate lyase in the central metabolism of mycobacteria the effect of growth rate on the in vivo fluxes was studied for the first time using ¹³C-metabolic flux analysis (MFA). Tracer experiments were performed with steady state chemostat cultures of BCG or M. tuberculosis supplied with ¹³C labeled glycerol or sodium bicarbonate. Through measurements of the ¹³C isotopomer labeling patterns in protein-derived amino acids and enzymatic activity assays we have identified the activity of a novel pathway for pyruvate dissimilation. We named this the GAS pathway because it utilizes the Glyoxylate shunt and Anapleurotic reactions for oxidation of pyruvate, and Succinyl CoA synthetase for the generation of succinyl CoA combined with a very low flux through the succinate--oxaloacetate segment of the tricarboxylic acid cycle. We confirm that M. tuberculosis can fix carbon from CO₂ into biomass. As the human host is abundant in CO₂ this finding requires further investigation in vivo as CO₂ fixation may provide a point of vulnerability that could be targeted with novel drugs. This study also provides a platform for further studies into the metabolism of M. tuberculosis using ¹³C-MFA.
Systems Biology has established numerous approaches for mechanistic modelling of molecular networks in the cell and a legacy of models. The current frontier is the integration of models expressed in different formalisms to address the multi-scale biological system organisation challenge. We present MUFINS software, implementing a unique set of approaches for multiformalism simulation of interaction networks. We extend the constraint-based modelling (CBM) framework by incorporation of linear inhibition constraints, enabling for the first time linear modelling of networks simultaneously describing gene regulation, signalling and whole-cell metabolism at steady state. We present a use case where a logical hypergraph model of a regulatory network is expressed by linear constraints and integrated with a Genome Scale Metabolic Network (GSMN) of mouse macrophage. We experimentally validate predictions, demonstrating application of our software in an iterative cycle of hypothesis generation, validation and model refinement. MUFINS incorporates an extended version of our Quasi Steady State Petri Net approach to integrate dynamic models with CBM, which we demonstrate through a dynamic model of cortisol signalling integrated with the human Recon2 GSMN and a model of nutrient dynamics in physiological compartments. Finally, we implement a number of methods for deriving metabolic states from ~omics data, including our new variant of the iMAT congruency approach. We compare our approach with iMAT through analysis of 262 individual tumour transcriptomes, recovering features of metabolic reprogramming in cancer. The software provides graphics user interface with network visualisation, which facilitates use by researchers who are not experienced in coding and mathematical modelling environments.
Enzymes at the phosphoenolpyruvate (PEP)–pyruvate–oxaloacetate or anaplerotic (ANA) node control the metabolic flux to glycolysis, gluconeogenesis, and anaplerosis. Here we used genetic, biochemical, and 13C isotopomer analysis to characterize the role of the enzymes at the ANA node in intracellular survival of the world's most successful bacterial pathogen, Mycobacterium tuberculosis (Mtb). We show that each of the four ANA enzymes, pyruvate carboxylase (PCA), PEP carboxykinase (PCK), malic enzyme (MEZ), and pyruvate phosphate dikinase (PPDK), performs a unique and essential metabolic function during the intracellular survival of Mtb. We show that in addition to PCK, intracellular Mtb requires PPDK as an alternative gateway into gluconeogenesis. Propionate and cholesterol detoxification was also identified as an essential function of PPDK revealing an unexpected role for the ANA node in the metabolism of these physiologically important intracellular substrates and highlighting this enzyme as a tuberculosis (TB)-specific drug target. We show that anaplerotic fixation of CO2 through the ANA node is essential for intracellular survival of Mtb and that Mtb possesses three enzymes (PCA, PCK, and MEZ) capable of fulfilling this function. In addition to providing a back-up role in anaplerosis we show that MEZ also has a role in lipid biosynthesis. MEZ knockout strains have an altered cell wall and were deficient in the initial entry into macrophages. This work reveals that the ANA node is a focal point for controlling the intracellular replication of Mtb, which goes beyond canonical gluconeogenesis and represents a promising target for designing novel anti-TB drugs.
Bovine tuberculosis is an important animal health problem and the predominant cause of zoonotic tuberculosis worldwide. It results in serious economic burden due to losses in productivity and the cost of control programmes. Control could be greatly improved by the introduction of an efficacious cattle vaccine but the most likely candidate, BCG, has several limitations including variable efficacy. Augmentation of BCG with a subunit vaccine booster has been shown to increase protection but the selection of antigens has hitherto been left largely to serendipity. In the present study, we take a rational approach to identify the protective antigens of BCG, selecting a BCG transposon mutant library in naïve and BCG-vaccinated cattle. Ten mutants had increased relative survival in vaccinated compared to naïve cattle, consistent with loss of protective antigen targets making the mutants less visible to the BCG immune response. The immunogenicity of three putative protective antigens, BCG_0116, BCG_0205 (YrbE1B) and BCG_1448 (PPE20) was investigated using peptide pools and PBMCs from BCG vaccinated cattle. BCG vaccination induced PBMC to release elevated levels of IP10, IL-17a and IL-10 in response to all three antigens. Taken together, the data supports the further study of these antigens for use in subunit vaccines.