Professor Dan Horton
Academic and research departments
School of Veterinary Medicine, Faculty of Health and Medical Sciences.About
Biography
Dan graduated as a veterinarian from the University of Cambridge, UK, with an intercalated MA in zoology in 2002. After a period in mixed and second opinion exotic animal veterinary practice he completed an MSc in Wild Animal Health in London in 2005. He then undertook a PhD working at Cambridge University, the APHA and CDC Atlanta USA, on zoonotic viral diseases of wildlife.
He completed his PhD in 2009 and joined the Virology Department at APHA Weybridge, undertaking surveillance and research programs for viral diseases of wildlife. Since February 2014 he has been at the School of Veterinary Medicine as a Lecturer, promoted to Senior Lecturer, Reader and then Professor of Veterinary Virology, continuing research into zoonotic viral diseases and teaching at undergraduate and postgraduate level. He is Associate Dean Research and Innovation for the Faculty of Health and Medical Sciences, an elected representative of Senate on the University Council, an Editor for PloS Neglected Tropical Diseases and a Diplomate of the European College of Zoological Medicine.
Areas of specialism
University roles and responsibilities
- Associate Dean, Research and Innovation (Faculty of Health and Medical Sciences)
- Senate Representative on University Council
My qualifications
News
In the media
ResearchResearch interests
Research in my group is focussed on reducing the animal and human health burden caused by diseases that cross species barriers.
Approaches include fundamental work on the evolution of viral pathogens, through to the applied aspects of control policy in current national and international rabies projects. An area of particular importance is the wildlife-livestock-human interface and its role in the emergence of pathogens. My previous research has given me a multifaceted perspective on this, including work characterising novel BSL3 pathogens in vitro and in-vivo, as well as planning, undertaking and interpreting data from surveillance programs for zoonotic pathogens in wildlife and domestic animals. Working in this area of viral zoonoses gives the one-health concept real meaning: in order to assess the risk posed by zoonotic infections you need to study the reservoir and the ecological niche that reservoir occupies, as well as the biology and behaviour of potential spill-over hosts.
An additional and allied area of interest is building veterinary and laboratory capacity internationally with obvious benefits for global biosecurity. This interest has been developed through several international collaborations on rabies, where I have combined technical assistance with research on the epidemiology of rabies.
Current work includes:
- OneHealth European Joint Programme (OneHealth EJP). A landmark European consortium project including 44 partners from 18 countries, taking a OneHealth approach to the challenges of foodborne zoonoses, antimicrobial resistance and emerging threats. Including:
- Co-Leader of the Joint Integrative Project COVRIN: One Health research integration on SARS-CoV-2 emergence, risk assessment and preparedness. Researchers Emma Taylor, Guido Cordoni
- Joint Research Projects:
- MAD-VIR Metagenomic Array Detection of emerging Viruses in EU, Post doctoral researcher Dr Elizabeth Royall
- TELE-VIR, a Point of Incidence toolbox for emerging virus threats Dr Carlijn Bogaardt, Dr Guido Cordoni
- MoMIR-PPC: Monitoring the gut microbiota and immune response to predict, prevent and control zoonoses in humans and livestock in order to minimize the use of antimicrobials. Post doctoral researcher: Dr Guido Cordoni
- Efficient risk management of One Health approaches: a portfolio analytic suite applied to rabies. Combining economic and disease dynamic models to improve decision making in disease control: Collaborators Dr Joaquin Prada, Dr Victor Del Rio Vilas, Dr Duncan Robertson (University of Loughborough) and US CDC. Postgraduate Researcher: Emma Taylor
- PRAGMATIST TRIP- The Road to Implementation. Validation and improvement of a vaccine optimisation tool for Foot and Mouth Disease with The Pirbright Institute and EU-FMD
- Exploiting the full potential of rapid diagnostic test technology and mobile connectivity for improved animal disease detection and control- with iSense consortium
- Poultry microbiome as a predictor of health & productivity : British Egg Marketing Board Pump Priming Award
- Ecological and evolutionary dynamics of the zoonotic pool: Collaboration with Dr Simon Anthony, Columbia USA and Dr Karin Darpel, Pirbright Institute. PhD candidate: Isamara Navarrete-Macias
- Improving prediction of zoonotic pathogen emergence and epidemics: Researcher Mobility Awards (University of Columbia)
- INSPIRE Award , supported by the Academy of Medical Sciences (on behalf of the School Research Culture Committee) Inspiring Veterinary Undergraduates to consider research careers
Recently completed projects:
- Understanding constraints and drivers acting on viruses that cross species barriers, using rabies virus as a model (Academy of Medical Sciences Springboard Award, supported by the Wellcome Trust), University Impact Acceleration fund
- Survival strategies of Rift Valley Fever Virus (PhD Student Sarah Lumley, collaborative supervisors Anthony Fooks, Nicholas Johnson (APHA), Roger Hewson (PHE) and Jolyon Medlock (PHE).
- Bats and Birds Research Network. British Council/Newton Fund research network with Pirbright Institute, USP Brazil, and APHA
Research interests
Research in my group is focussed on reducing the animal and human health burden caused by diseases that cross species barriers.
Approaches include fundamental work on the evolution of viral pathogens, through to the applied aspects of control policy in current national and international rabies projects. An area of particular importance is the wildlife-livestock-human interface and its role in the emergence of pathogens. My previous research has given me a multifaceted perspective on this, including work characterising novel BSL3 pathogens in vitro and in-vivo, as well as planning, undertaking and interpreting data from surveillance programs for zoonotic pathogens in wildlife and domestic animals. Working in this area of viral zoonoses gives the one-health concept real meaning: in order to assess the risk posed by zoonotic infections you need to study the reservoir and the ecological niche that reservoir occupies, as well as the biology and behaviour of potential spill-over hosts.
An additional and allied area of interest is building veterinary and laboratory capacity internationally with obvious benefits for global biosecurity. This interest has been developed through several international collaborations on rabies, where I have combined technical assistance with research on the epidemiology of rabies.
Current work includes:
- OneHealth European Joint Programme (OneHealth EJP). A landmark European consortium project including 44 partners from 18 countries, taking a OneHealth approach to the challenges of foodborne zoonoses, antimicrobial resistance and emerging threats. Including:
- Co-Leader of the Joint Integrative Project COVRIN: One Health research integration on SARS-CoV-2 emergence, risk assessment and preparedness. Researchers Emma Taylor, Guido Cordoni
- Joint Research Projects:
- MAD-VIR Metagenomic Array Detection of emerging Viruses in EU, Post doctoral researcher Dr Elizabeth Royall
- TELE-VIR, a Point of Incidence toolbox for emerging virus threats Dr Carlijn Bogaardt, Dr Guido Cordoni
- MoMIR-PPC: Monitoring the gut microbiota and immune response to predict, prevent and control zoonoses in humans and livestock in order to minimize the use of antimicrobials. Post doctoral researcher: Dr Guido Cordoni
- Efficient risk management of One Health approaches: a portfolio analytic suite applied to rabies. Combining economic and disease dynamic models to improve decision making in disease control: Collaborators Dr Joaquin Prada, Dr Victor Del Rio Vilas, Dr Duncan Robertson (University of Loughborough) and US CDC. Postgraduate Researcher: Emma Taylor
- PRAGMATIST TRIP- The Road to Implementation. Validation and improvement of a vaccine optimisation tool for Foot and Mouth Disease with The Pirbright Institute and EU-FMD
- Exploiting the full potential of rapid diagnostic test technology and mobile connectivity for improved animal disease detection and control- with iSense consortium
- Poultry microbiome as a predictor of health & productivity : British Egg Marketing Board Pump Priming Award
- Ecological and evolutionary dynamics of the zoonotic pool: Collaboration with Dr Simon Anthony, Columbia USA and Dr Karin Darpel, Pirbright Institute. PhD candidate: Isamara Navarrete-Macias
- Improving prediction of zoonotic pathogen emergence and epidemics: Researcher Mobility Awards (University of Columbia)
- INSPIRE Award , supported by the Academy of Medical Sciences (on behalf of the School Research Culture Committee) Inspiring Veterinary Undergraduates to consider research careers
Recently completed projects:
- Understanding constraints and drivers acting on viruses that cross species barriers, using rabies virus as a model (Academy of Medical Sciences Springboard Award, supported by the Wellcome Trust), University Impact Acceleration fund
- Survival strategies of Rift Valley Fever Virus (PhD Student Sarah Lumley, collaborative supervisors Anthony Fooks, Nicholas Johnson (APHA), Roger Hewson (PHE) and Jolyon Medlock (PHE).
- Bats and Birds Research Network. British Council/Newton Fund research network with Pirbright Institute, USP Brazil, and APHA
Supervision
Postgraduate research supervision
Current postgraduate research students
Emma Taylor: Efficient risk management of One Health approaches: a portfolio analytic suite applied to rabies. Collaborators CDC, Prada Modelling Lab
Philippa Hollinghurst: Evolution of Rift Valley Fever Virus. (collaboration with the mosquito immunology group, Pirbright Institute)
Rhiannon Moody: “Not all viruses come alone” - the impact of Orbivirus co-infections on bluetongue virus reassortment and diversity (collaboration with the Orbivirus research group, Pirbright Institute)
Abdelaziz Yassin: Development of serological tools to support the control of foot-and-mouth disease in Egypt. (collaboration with the World Reference Laboratory for FMDV at the Pirbright Institute)
Megan Golding: Evolution and infection dynamics of European Bat Lyssavirus 1 and its host Eptesicus serotinus in the UK. (Collaboration with the Virology Department, Animal and Plant Health Agency)
Natasha Edwards: Forecasting the next panzootic foot-and-mouth disease lineage: informing virus fitness from in vitro studies and genomic data (collaboration with World Reference Laboratory for FMDV, Pirbright Institute)
Jennifer Rokhsar (co-supervisor)
Completed post graduate research students:
Dr Sarah Lumley (DSTL, UK)
Dr Isamara Navarrete Macias (UC Davis, USA)
Dr Muneeb Islam (Department of Microbiology at Abdul Wali Khan University, Pakistan)
Teaching
- Module coordinator MSc Veterinary Microbiology Module 10: 'Systems diseases of animals: CNS and Skin'
- Contributions to Veterinary Medicine and Science Foundations of Disease modules (Level 5), Clinical practice (Level 6)
- MSc Medical Microbiology (Level 7)
- Biomedical Science Vet Public Health and Animal Infectious Diseases
- Problem based learning (Levels 4 and 5) and OSCEs
- Programme Leader MSc Veterinary Microbiology 2016-2019
Publications
Neglected tropical diseases (NTDs) disproportionately affect the poorest populations. To meet the 2030 World Health Organization (WHO) roadmap targets, integrated and cross-cutting approaches are recommended to streamline programmatic operations across NTDs. Integration is not a new concept. But given recent policy shifts, disruptions to programmes and funding constraints, it is increasingly important. We consider integration as opportunities for coordination or collaboration across NTDs. Research is needed to identify the criteria and requirements necessary for successful programme integration, assess barriers, and determine the relative importance of each criterion to identify potential disease pairings. We applied Multi-Criteria Decision Analysis (MCDA) methodologies to gather expert and stakeholder insights on integrated control programmes for NTDs. During a facilitated workshop, participants discussed their interpretations of the terms 'programmatic integration' and 'cross-cutting' in relation to integrating NTDs programmes. 11 criteria for integration were identified and weighed by participants. Using WHO Roadmap baseline values, pairwise disease combinations were assessed by multiplying criterion weightings with disease scores, generating a priority matrix. Workshop participants weighted community engagement and common vectors and transmission routes as the most important of the criteria. Three disease combinations with the highest potential for integration were identified, Dengue and Chikungunya, Taeniasis & Cysticercosis and Echinococcosis, and Trachoma and Lymphatic Filariasis. The workshop outcomes provide valuable insights into key factors for integrating NTD control programmes and highlight potential disease pairings for further exploration. While some disease matches were expected, others were less obvious. The highest-scoring combinations should now be further evaluated for integration potential.
Rabies was first reported in ancient Iraqi civilizations, yet it remains a poorly quantified and important public health threat in the region. Efforts to control rabies in Iraq including dog population control, and vaccination of livestock and dogs, have increased since 2010. Officially reported data on human rabies, dog bites, and animal rabies cases between 2012 and 2017 are analysed here to assess the effect of existing control efforts, to inform future strategies, and to highlight gaps in surveillance and reporting. The results of molecular characterization of 32 viruses from animal cases from throughout Iraq are presented, to improve the understanding of rabies dynamics in the animal reservoir. Although annual numbers of reported human cases were lower in the period between 2012 and 2017 than prior to 2010, human cases continue. There was a distinct gender and age bias among human cases with nine cases in males for every one female and twice as many cases in children than adults. Spatial clustering analysis and phylogenetic evidence suggests rabies is endemic throughout the country, with no regional variation in risk, but better surveillance and reporting is required to underpin control strategies.
Rabies is a feared global disease, largely due to the extreme case-fatality rate, which approaches 100% once a patient becomes ill [4]. The true burden of disease remains undefined, but rabies causes tens of thousands of human deaths annually and is responsible for major economic losses, including dog vaccination costs in excess of US$130 million, and US$512 million in livestock deaths in rabies-endemic countries. After exposure, rabies virus causes an acute, progressive fatal encephalitis, which is prevented with timely post-exposure prophylaxis (PEP). Pre-immunisation in populations at risk, using modern, safe and efficacious cell culture vaccines, also prevents rabies. Where the disease burden is greatest, vaccines are not always readily available, and they can be prohibitively expensive. Rabies cases in vaccinated individuals are becoming increasingly recognised [6]. The challenge of ineffective vaccines is complex and multifactorial but could be part of an apparent growing global trend for counterfeit vaccines [2]. The World Health Organization (WHO) defines counterfeit medicines, including vaccines, as products containing either incorrect, little or no active ingredient, which are marketed and sold as authentic medicines. Such products are illegal if the contents have been altered deliberately and the authorisation forged. The increasing availability and administration of counterfeit rabies vaccines, especially for PEP, may result in otherwise preventable deaths. In 2015, the WHO, the World Organisation for Animal Health, and the Food and Agriculture Organization of the United Nations, in partnership with the Global Alliance for Rabies Control, proposed the global goal of zero human deaths as a result of dog-mediated rabies, by 2030. This goal is supported by a strategic plan, aiming to save lives, reduce costs, and enable clear communications on the validation of canine rabies elimination in endemic countries.
The global 2030 goal set by the World Organization for Animal Health (WOAH), the World Health Organization (WHO), and the Food and Agriculture Organization (FAO), to eliminate dog-mediated human rabies deaths, has undeniably been a catalyst for many countries to re-assess existing dog rabies control programmes. Additionally, the 2030 agenda for Sustainable Development includes a blueprint for global targets which will benefit both people and secure the health of the planet. Rabies is acknowledged as a disease of poverty, but the connections between economic development and rabies control and elimination are poorly quantified yet, critical evidence for planning and prioritisation. We have developed multiple generalised linear models, to model the relationship between health care access, poverty, and death rate as a result of rabies, with separate indicators that can be used at country-level; total Gross Domestic Product (GDP), and current health expenditure as a percentage of the total gross domestic product (% GDP) as an indicator of economic growth; and a metric of poverty assessing the extent and intensity of deprivation experienced at the individual level (Multidimensional Poverty Index, MPI). Notably there was no detectable relationship between GDP or current health expenditure (% GDP) and death rate from rabies. However, MPI showed statistically significant relationships with per capita rabies deaths and the probability of receiving lifesaving post exposure prophylaxis. We highlight that those most at risk of not being treated, and dying due to rabies, live in communities experiencing health care inequalities, readily measured through poverty indicators. These data demonstrate that economic growth alone, may not be enough to meet the 2030 goal. Indeed, other strategies such as targeting vulnerable populations and responsible pet ownership are also needed in addition to economic investment.
The Middle East, Eastern Europe, Central Asia and North Africa Rabies Control Network (MERACON), is built upon the achievements of the Middle East and Eastern Europe Rabies Expert Bureau (MEEREB). MERACON aims to foster collaboration among Member States (MS) and develop shared regional objectives, building momentum towards dog-mediated rabies control and elimination. Here we assess the epidemiology of rabies and preparedness in twelve participating MS, using case and rabies capacity data for 2017, and compare our findings with previous published reports and a predictive burden model. Across MS, the number of reported cases of dog rabies per 100,000 dog population and the number of reported human deaths per 100,000 population as a result of dog-mediated rabies appeared weakly associated. Compared to 2014 there has been a decrease in the number of reported human cases in five of the twelve MS, three MS reported an increase, two MS continued to report zero cases, and the remaining two MS were not listed in the 2014 study and therefore no comparison could be drawn. Vaccination coverage in dogs has increased since 2014 in half (4/8) of the MS where data are available. Most importantly, it is evident that there is a need for improved data collection, sharing and reporting at both the national and international levels. With the formation of the MERACON network, MS will be able to align with international best practices, while also fostering international support with other MS and international organisations.
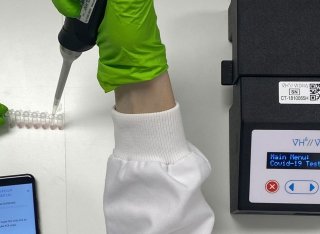
In the published article, there was an error in the Article title. Instead of “VIDIIA Hunter diagnostic platform: a low-cost, smartphone connected, artificial intelligence-assisted COVID-19 rapid diagnostics approved for medical use in the UK,” it should be “VIDIIA Hunter: a low-cost, smartphone connected, artificial intelligence-assisted COVID-19 rapid diagnostic platform approved for medical use in the UK”. The authors apologize for this error and state that this does not change the scientific conclusions of the article in any way. The original article has been updated.
Introduction: Accurate and rapid diagnostics paired with effective tracking and tracing systems are key to halting the spread of infectious diseases, limiting the emergence of new variants and to monitor vaccine efficacy. The current gold standard test (RT-qPCR) for COVID-19 is highly accurate and sensitive, but is time-consuming, and requires expensive specialised, lab-based equipment. Methods: Herein, we report on the development of a SARS-CoV-2 (COVID-19) rapid and inexpensive diagnostic platform that relies on a reverse-transcription loop-mediated isothermal amplification (RT-LAMP) assay and a portable smart diagnostic device. Automated image acquisition and an Artificial Intelligence (AI) deep learning model embedded in the Virus Hunter 6 (VH6) device allow to remove any subjectivity in the interpretation of results. The VH6 device is also linked to a smartphone companion application that registers patients for swab collection and manages the entire process, thus ensuring tests are traced and data securely stored. Results: Our designed AI-implemented diagnostic platform recognises the nucleocapsid protein gene of SARS-CoV-2 with high analytical sensitivity and specificity. A total of 752 NHS patient samples, 367 confirmed positives for coronavirus disease (COVID-19) and 385 negatives, were used for the development and validation of the test and the AI-assisted platform. The smart diagnostic platform was then used to test 150 positive clinical samples covering a dynamic range of clinically meaningful viral loads and 250 negative samples. When compared to RT-qPCR, our AI-assisted diagnostics platform was shown to be reliable, highly specific (100%) and sensitive (98–100% depending on viral load) with a limit of detection of 1.4 copies of RNA per µL in 30 min. Using this data, our CE-IVD and MHRA approved test and associated diagnostic platform has been approved for medical use in the United Kingdom under the UK Health Security Agency’s Medical Devices (Coronavirus Test Device Approvals, CTDA) Regulations 2022. Laboratory and in-silico data presented here also indicates that the VIDIIA diagnostic platform is able to detect the main variants of concern in the United Kingdom (September 2023). Discussion: This system could provide an efficient, time and cost-effective platform to diagnose SARS-CoV-2 and other infectious diseases in resource-limited settings.
Pigs infected with Salmonella may excrete large amounts of Salmonella, increasing the risk of spread of this pathogen in the food chain. Identifying Salmonella high shedder pigs is therefore required to mitigate this risk. We analyzed immune-associated markers and composition of the gut microbiota in specific-pathogen-free pigs presenting different shedding levels after an oral infection with Salmonella. Immune response was studied through total blood cell counts, production of anti-Salmonella antibodies and cytokines, and gene expression quantification. Total Salmonella shedding for each pig was estimated and hierarchical clustering was used to cluster pigs into high, intermediate, and low shedders. Gut microbiota compositions were assessed using 16S rRNA microbial community profiling. Comparisons were made between control and inoculated pigs, then between high and low shedders pigs. Prior to infection, high shedders had similar immunological profiles compared to low shedders. As soon as 1 day postinoculation (dpi), significant differences on the cytokine production level and on the expression level of several host genes related to a proinflammatory response were observed between high and low shedders. Infection with Salmonella induced an early and profound remodeling of the immune response in all pigs, but the intensity of the response was stronger in high shedders. In contrast, low shedders seroconverted earlier than high shedders. Just after induction of the proinflammatory response (at 2 dpi), some taxa of the fecal microbiota were specific to the shedding phenotypes. This was related to the enrichment of several functional pathways related to anaerobic respiration in high shedders. In conclusion, our data show that the immune response to Salmonella modifies the fecal microbiota and subsequently could be responsible for shedding phenotypes. Influencing the gut microbiota and reducing intestinal inflammation could be a strategy for preventing Salmonella high shedding in livestock. Salmonellosis remains the most frequent human foodborne zoonosis after campylobacteriosis and pork meat is considered one of the major sources of human foodborne infections. At the farm, host heterogeneity in pig infection is problematic. High Salmonella shedders contribute more significantly to the spread of this foodborne pathogen in the food chain. The identification of predictive biomarkers for high shedders could help to control Salmonella in pigs. The purpose of the present study was to investigate why some pigs become super shedders and others low shedders. We thus investigated the differences in the fecal microbial composition and the immune response in orally infected pigs presenting different Salmonella shedding patterns. Our data show that the proinflammatory response induced by Typhimurium at 1 dpi could be responsible for the modification of the fecal microbiota composition and functions observed mainly at 2 and 3 dpi and to the low and super shedder phenotypes.
The sequencing of viral genomes provides important data for the prevention and control of foot-and-mouth disease (FMD) outbreaks. Sequence data can be used for strain identification, outbreak tracing, and aiding the selection of the most appropriate vaccine for the circulating strains. At present, sequencing of FMD virus (FMDV) relies upon the time-consuming transport of samples to well-resourced laboratories. The Oxford Nanopore Technologies' MinION portable sequencer has the potential to allow sequencing in remote, decentralised laboratories closer to the outbreak location. In this study, we investigated the utility of the MinION to generate sequence data of sufficient quantity and quality for the characterisation of FMDV serotypes O, A, Asia 1. Prior to sequencing, a universal two-step RT-PCR was used to amplify parts of the 5′UTR, as well as the leader, capsid and parts of the 2A encoding regions of FMDV RNA extracted from three sample matrices: cell culture supernatant, tongue epithelial suspension and oral swabs. The resulting consensus sequences were compared with reference sequences generated on the Illumina MiSeq platform. Consensus sequences with an accuracy of 100% were achieved within 10 and 30 min from the start of the sequencing run when using RNA extracted from cell culture supernatants and tongue epithelial suspensions, respectively. In contrast, sequencing from swabs required up to 2.5 h. Together these results demonstrated that the MinION sequencer can be used to accurately and rapidly characterise serotypes A, O, and Asia 1 of FMDV using amplicons amplified from a variety of different sample matrices.
Passive surveillance for lyssaviruses in UK bats has been ongoing since 1987 and has identified 13 cases of EBLV-2 from a single species; Myotis daubentonii. No other lyssavirus species has been detected. Between 2005 and 2015, 10 656 bats were submitted, representing 18 species, creating a spatially and temporally uneven sample of British bat fauna. Uniquely, three UK cases originate from a roost at Stokesay Castle in Shropshire, England, where daily checks for grounded and dead bats are undertaken and bat carcasses have been submitted for testing since 2007. Twenty per cent of Daubenton's bats submitted from Stokesay Castle since surveillance began, have tested positive for EBLV-2. Phylogenetic analysis reveals geographical clustering of UK viruses. Isolates from Stokesay Castle are more closely related to one another than to viruses from other regions. Daubenton's bats from Stokesay Castle represent a unique opportunity to study a natural population that appears to maintain EBLV-2 infection and may represent endemic infection at this site. Although the risk to public health from EBLV-2 is low, consequences of infection are severe and effective communication on the need for prompt post-exposure prophylaxis for anyone that has been bitten by a bat is essential.
The Middle East is a culturally and politically diverse region at the gateway between Europe, Africa and Asia. Spatial dynamics of the fatal zoonotic disease rabies among countries of the Middle East and surrounding regions is poorly understood. An improved understanding of virus distribution is necessary to direct control methods. Previous studies have suggested regular trans-boundary movement, but have been unable to infer direction. Here we address these issues, by investigating the evolution of 183 rabies virus isolates collected from over 20 countries between 1972 and 2014. We have undertaken a discrete phylogeographic analysis on a subset of 139 samples to infer where and when movements of rabies have occurred. We provide evidence for four genetically distinct clades with separate origins currently circulating in the Middle East and surrounding countries. Introductions of these viruses have been followed by regular and multidirectional trans-boundary movements in some parts of the region, but relative isolation in others. There is evidence for minimal regular incursion of rabies from Central and Eastern Asia. These data support current initiatives for regional collaboration that are essential for rabies elimination.
Background With the advent of Next Generation Sequencing (NGS) technologies, the ability to generate large amounts of sequence data has revolutionized the genomics field. Most RNA viruses have relatively small genomes in comparison to other organisms and as such, would appear to be an obvious success story for the use of NGS technologies. However, due to the relatively low abundance of viral RNA in relation to host RNA, RNA viruses have proved relatively difficult to sequence using NGS technologies. Here we detail a simple, robust methodology, without the use of ultra-centrifugation, filtration or viral enrichment protocols, to prepare RNA from diagnostic clinical tissue samples, cell monolayers and tissue culture supernatant, for subsequent sequencing on the Roche 454 platform. Results As representative RNA viruses, full genome sequence was successfully obtained from known lyssaviruses belonging to recognized species and a novel lyssavirus species using these protocols and assembling the reads using de novo algorithms. Furthermore, genome sequences were generated from considerably less than 200 ng RNA, indicating that manufacturers’ minimum template guidance is conservative. In addition to obtaining genome consensus sequence, a high proportion of SNPs (Single Nucleotide Polymorphisms) were identified in the majority of samples analyzed. Conclusions The approaches reported clearly facilitate successful full genome lyssavirus sequencing and can be universally applied to discovering and obtaining consensus genome sequences of RNA viruses from a variety of sources.
Lyssaviruses (family Rhabdoviridae) constitute one of the most important groups of viral zoonoses globally. All lyssaviruses cause the disease rabies, an acute progressive encephalitis for which, once symptoms occur, there is no effective cure. Currently available vaccines are highly protective against the predominantly circulating lyssavirus species. Using next-generation sequencing technologies, we have obtained the whole-genome sequence for a novel lyssavirus, Ikoma lyssavirus (IKOV), isolated from an African civet in Tanzania displaying clinical signs of rabies. Genetically, this virus is the most divergent within the genus Lyssavirus. Characterization of the genome will help to improve our understanding of lyssavirus diversity and enable investigation into vaccine-induced immunity and protection.
Vampire bat-transmitted rabies has recently become the leading cause of rabies mortality in both humans and livestock in Latin America. Evaluating risk of transmission from bats to other animal species has thus become a priority in the region. An integrated bat-rabies dynamic modeling framework quantifying spillover risk to cattle farms was developed. The model is spatially explicit and is calibrated to the state of São Paulo, using real roost and farm locations. Roost and farm characteristics, as well as environmental data through an ecological niche model, are used to modulate rabies transmission. Interventions aimed at reducing risk in roosts (such as bat culling or vaccination) and in farms (cattle vaccination) were considered as control strategies. Both interventions significantly reduce the number of outbreaks in farms and disease spread (based on distance from source), with control in roosts being a significantly better intervention. High-risk areas were also identified, which can support ongoing programs, leading to more effective control interventions.
Control of rabies requires a consistent supply of dependable resources, constructive cooperation between veterinary and public health authorities, and systematic surveillance. These are challenging in any circumstances, but particularly during conflict. Here we describe available human rabies surveillance data from Iraq, results of renewed sampling for rabies in animals, and the first genetic characterisation of circulating rabies strains from Iraq. Human rabies is notifiable, with reported cases increasing since 2003, and a marked increase in Baghdad between 2009 and 2010. These changes coincide with increasing numbers of reported dog bites. There is no laboratory confirmation of disease or virus characterisation and no systematic surveillance for rabies in animals. To address these issues, brain samples were collected from domestic animals in the greater Baghdad region and tested for rabies. Three of 40 brain samples were positive using the fluorescent antibody test and hemi-nested RT-PCR for rabies virus (RABV). Bayesian phylogenetic analysis using partial nucleoprotein gene sequences derived from the samples demonstrated the viruses belong to a single virus variant and share a common ancestor with viruses from neighbouring countries, 22 (95% HPD 14-32) years ago. These include countries lying to the west, north and east of Iraq, some of which also have other virus variants circulating concurrently. These results suggest possible multiple introductions of rabies into the Middle East, and regular trans-boundary movement of disease. Although 4000 years have passed since the original description of disease consistent with rabies, animals and humans are still dying of this preventable and neglected zoonosis.
The successful prevention, control, and elimination of dog-mediated rabies is challenging due to insufficient resource availability and inadequate placement. An integrated dog bite case management (IBCM) system plus dog vaccination can help address these challenges. Based on data from the IBCM system in Haiti, we conducted a cost-effectiveness evaluation of a newly established IBCM system plus sustained vaccination and compared it with 1) a no bite-case management (NBCM) and 2) a non–risk-based (NRB) program, where bite victims presenting at a health clinic would receive post-exposure prophylaxis regardless of risk assessment. We also provide cost-effectiveness guidance for an ongoing IBCM system and for sub-optimal dog vaccination coverages, considering that not all cost-effective interventions are affordable. Cost-effectiveness outcomes included average cost per human death averted (USD/death averted) and per life-year gained (LYG). The analysis used a governmental perspective. Considering a sustained 5-year implementation with 70% dog vaccination coverage, IBCM had a lower average cost per death averted (IBCM: $7,528, NBCM: $7,797, NRB: $15,244) and cost per LYG (IBCM: $152, NBCM: $158, NRB: $308) than NBCM and NRB programs. As sensitivity analysis, we estimated cost-effectiveness for alternative scenarios with lower dog-vaccination coverages (30%, 55%) and lower implementation costs. Our results suggest that better health and cost-effectiveness outcomes are achieved with the continued implementation of an IBCM program ($118 per life-year saved) compared with a newly established IBCM program ($152 per life-year saved). Our results suggest that IBCM is more cost-effective than non-integrated programs to eliminate dog-mediated human rabies.
Rabies is a fatal neurologic disease caused by lyssavirus infection. Bats are important natural reservoir hosts of various lyssaviruses that can be transmitted to people. The epidemiology and pathogenesis of rabies in bats are poorly understood, making it difficult to prevent zoonotic transmission. To further our understanding of lyssavirus pathogenesis in a natural bat host, an experimental model using straw-colored fruit bats (Eidolon helvum) and Lagos bat virus, an endemic lyssavirus in this species, was developed. To determine the lowest viral dose resulting in 100% productive infection, bats in five groups (four bats per group) were inoculated intramuscularly with one of five doses, ranging from 100.1 to 104.1 median tissue culture infectious dose (TCID50). More bats died due to the development of rabies after the middle dose (102.1 TCID50, 4/4 bats) than after lower (101.1, 2/4; 101.1, 2/4) or higher (103.1, 2/4; 104.1, 2/4) doses of virus. In the two highest dose groups, 4/8 bats developed rabies. Of those bats that remained healthy 3/4 bats seroconverted, suggesting that high antigen loads can trigger a strong immune response that abrogates a productive infection. In contrast, in the two lowest dose groups, 3/8 bats developed rabies, 1/8 remained healthy and seroconverted and 4/8 bats remained healthy and did not seroconvert, suggesting these doses are too low to reliably induce infection. The main lesion in all clinically affected bats was meningoencephalitis associated with lyssavirus-positive neurons. Lyssavirus antigen was detected in tongue epithelium (5/11 infected bats) rather than in salivary gland epithelium (0/11), suggesting viral excretion via the tongue. Thus, intramuscular inoculation of 102.1 TCID50 of Lagos bat virus into straw-colored fruit bats is a suitable model for lyssavirus associated bat rabies in a natural reservoir host, and can help with the investigation of lyssavirus infection dynamics in bats.
Background: The chicken is the most abundant food animal in the world. However, despite its importance, the chicken gut microbiome remains largely undefined. Here, we exploit culture-independent and culture-dependent approaches to reveal extensive taxonomic diversity within this complex microbial community. Results: We performed metagenomic sequencing of fifty chicken faecal samples from two breeds and analysed these, alongside all (n = 582) relevant publicly available chicken metagenomes, to cluster over 20 million non-redundant genes and to construct over 5,500 metagenome-assembled bacterial genomes. In addition, we recovered nearly 600 bacteriophage genomes. This represents the most comprehensive view of taxonomic diversity within the chicken gut microbiome to date, encompassing hundreds of novel candidate bacterial genera and species. To provide a stable, clear and memorable nomenclature for novel species, we devised a scalable combinatorial system for the creation of hundreds of well-formed Latin binomials. We cultured and genome-sequenced bacterial isolates from chicken faeces, documenting over forty novel species, together with three species from the genus Escherichia, including the newly named species Escherichia whittamii. Conclusions: Our metagenomic and culture-based analyses provide new insights into the bacterial, archaeal and bacteriophage components of the chicken gut microbiome. The resulting datasets expand the known diversity of the chicken gut microbiome and provide a key resource for future high-resolution taxonomic and functional studies on the chicken gut microbiome.
Antigen banks have been established to supply foot-and-mouth disease virus (FMDV) vaccines at short notice to respond to incursions or upsurges in cases of FMDV infection. Multiple vaccine strains are needed to protect against specific FMDV lineages that circulate within six viral serotypes that are unevenly distributed across the world. The optimal selection of distinct antigens held in a bank must carefully balance the desire to cover these risks with the costs of purchasing and maintaining vaccine antigens. PRAGMATIST is a semi-quantitative FMD vaccine strain selection tool combining three strands of evidence: ( 1 ) estimates of the risk of incursion from specific areas (source area score); ( 2 ) estimates of the relative prevalence of FMD viral lineages in each specific area (lineage distribution score); and ( 3 ) effectiveness of each vaccine against specific FMDV lineages based on laboratory vaccine matching tests (vaccine coverage score). The output is a vaccine score, which identifies vaccine strains that best address the threats, and consequently which are the highest priority for inclusion in vaccine antigen banks. In this paper, data used to populate PRAGMATIST are described, including the results from expert elicitations regarding FMD risk and viral lineage circulation, while vaccine coverage data is provided from vaccine matching tests performed at the WRLFMD between 2011 and 2021 ( n = 2,150). These data were tailored to working examples for three hypothetical vaccine antigen bank perspectives (Europe, North America, and Australia). The results highlight the variation in the vaccine antigens required for storage in these different regions, dependent on risk. While the tool outputs are largely robust to uncertainty in the input parameters, variation in vaccine coverage score had the most noticeable impact on the estimated risk covered by each vaccine, particularly for vaccines that provide substantial risk coverage across several lineages.
In 2009, a novel lyssavirus (subsequently named Ikoma lyssavirus, IKOV) was detected in the brain of an African civet (Civettictis civetta) with clinical rabies in the Serengeti National Park of Tanzania. The degree of nucleotide divergence between the genome of IKOV and those of other lyssaviruses predicted antigenic distinction from, and lack of protection provided by, available rabies vaccines. In addition, the index case was considered likely to be an incidental spillover event, and therefore the true reservoir of IKOV remained to be identified. The advent of sensitive molecular techniques has led to a rapid increase in the discovery of novel viruses. Detecting viral sequence alone, however, only allows for prediction of phenotypic characteristics and not their measurement. In the present study we describe the in vitro and in vivo characterization of IKOV, demonstrating that it is (1) pathogenic by peripheral inoculation in an animal model, (2) antigenically distinct from current rabies vaccine strains and (3) poorly neutralized by sera from humans and animals immunized against rabies. In a laboratory mouse model, no protection was elicited by a licensed rabies vaccine. We also investigated the role of bats as reservoirs of IKOV. We found no evidence for infection among 483 individuals of at least 13 bat species sampled across sites in the Serengeti and Southern Kenya.
Overview One Health is an increasingly popular approach used to tackle complex health problems. The One Health concept recognizes that human health is tightly connected to the health of animals and the environment. Although the related fields are now more aware of the benefits of collaborative working, the full benefits have not yet been realized as research efforts are often focussed on just one of these health domains. To address regional and global issues such as foodborne zoonoses (FBZ), antimicrobial resistance (AMR) and emerging infectious threats (ET), there must be transdisciplinary collaboration between the health domains, in addition to active dialogue between scientists and international policy makers. This editorial introduces the One Health European Joint Programme (OHEJP) as an example of a One Health initiative. Zoonoses, AMR and their global burden Zoonoses are infectious diseases that can be transmitted directly or indirectly between humans and animals. Although the severity of zoonotic infections varies, their global impact is undisputable. The World Bank estimates that just six zoonotic disease outbreaks between 1997 and 2009 led to a global economic loss of US$ 80 billion [1]. This high cost is due to medical costs, loss of individual productivity and restrictions on trade and movement during outbreaks. Despite improvements in the management and treatment of zoonotic outbreaks, high disease burdens caused by zoonotic pathogens continue to be reported globally. These problems have been amply demonstrated recently by the SARS-CoV-2 pandemic. Although it is still too soon to fully assess the total economic and societal cost of this virus, recent publications, such as Nicola et al. [2] have begun to highlight just how widespread the impact of a truly global zoonotic disease can be. Alongside zoonoses, AMR is a growing international issue. The World Health Organization (WHO) has listed AMR as one of the ten greatest global health threats in 2019 [3]. AMR is defined as the ability of microorganisms to survive the effect of antimicrobial drugs, hindering not only our ability to treat infectious diseases, but also to perform medical procedures requiring prophylactic antibiotic administration. It has been predicted that by 2050, the number of deaths due to unresponsive infections will reach 10 million annually, with the associated costs being estimated at US$ 100 trillion [4]. Increased and inappropriate use of antimicrobials has contributed to the development and spread of AMR, which can be transmitted between humans, animals and the environment. The history of the ‘One Health’ concept The origins of One Health go as far back as 1855, when Rudolf Virchow founded comparative pathology, which could be seen as the origin of the One Health concept. Building upon this, Calvin W. Schwabe argued in the twentieth century against compartmentalization in medical research, using the term ‘One Medicine’. The term One Health was then popularized in 2004 by the Wildlife Conservation Society at a conference in New York [5], and its use has continued to evolve since then, fostering the revival of comparative medicine (Fig. 1, and reviewed in Gibbs [6]). One Health has now been adopted by the WHO [7], the Food and Agriculture Organization (FAO) [8] and the World Organization for Animal Health (OIE) [9].
Rabies is a fatal neurologic disease caused by lyssavirus infection. People are infected through contact with infected animals. The relative increase of human rabies acquired from bats calls for a better understanding of lyssavirus infections in their natural hosts. So far, there is no experimental model that mimics natural lyssavirus infection in the reservoir bat species. Lagos bat virus is a lyssavirus that is endemic in straw-colored fruit bats (Eidolon helvum) in Africa. Here we compared the susceptibility of these bats to three strains of Lagos bat virus (from Senegal, Nigeria, and Ghana) by intracranial inoculation. To allow comparison between strains, we ensured the same titer of virus was inoculated in the same location of the brain of each bat. All bats (n = 3 per strain) were infected, and developed neurological signs, and fatal meningoencephalitis with lyssavirus antigen expression in neurons. There were three main differences among the groups. First, time to death was substantially shorter in the Senegal and Ghana groups (4 to 6 days) than in the Nigeria group (8 days). Second, each virus strain produced a distinct clinical syndrome. Third, the spread of virus to peripheral tissues, tested by hemi-nested reverse transcriptase PCR, was frequent (3 of 3 bats) and widespread (8 to 10 tissues positive of 11 tissues examined) in the Ghana group, was frequent and less widespread in the Senegal group (3/3 bats, 3 to 6 tissues positive), and was rare and restricted in the Nigeria group (1/3 bats, 2 tissues positive). Centrifugal spread of virus from brain to tissue of excretion in the oral cavity is required to enable lyssavirus transmission. Therefore, the Senegal and Ghana strains seem most suitable for further pathogenesis, and for transmission, studies in the straw-colored fruit bat.
The current measures to control foot-and-mouth disease (FMD) include vaccination, movement control and slaughter of infected or susceptible animals. One of the difficulties in controlling FMD by vaccination arises due to the substantial diversity found among the seven serotypes of FMD virus (FMDV) and the strains within these serotypes. Therefore, vaccination using a single vaccine strain may not fully cross-protect against all strains within that serotype, and therefore selection of appropriate vaccines requires serological comparison of the field virus and potential vaccine viruses using relationship coefficients (r1 values). Limitations of this approach are that antigenic relationships among field viruses are not addressed, as comparisons are only with potential vaccine virus. Furthermore, inherent variation among vaccine sera may impair reproducibility of one-way relationship scores. Here, we used antigenic cartography to quantify and visualize the antigenic relationships among FMD serotype A viruses, aiming to improve the understanding of FMDV antigenic evolution and the scope and reliability of vaccine matching. Our results suggest that predicting antigenic difference using genetic sequence alone or by geographical location is not currently reliable. We found co-circulating lineages in one region that were genetically similar but antigenically distinct. Nevertheless, by comparing antigenic distances measured from the antigenic maps with the full capsid (P1) sequence, we identified a specific amino acid substitution associated with an antigenic mismatch among field viruses and a commonly used prototype vaccine strain, A22/IRQ/24/64.
Background Neglected tropical diseases (NTDs) disproportionately affect populations living in resource-limited settings. In the Amazon basin, substantial numbers of NTDs are zoonotic, transmitted by vertebrate (dogs, bats, snakes) and invertebrate species (sand flies and triatomine insects). However, no dedicated consortia exist to find commonalities in the risk factors for or mitigations against bite-associated NTDs such as rabies, snake envenoming, Chagas disease and leishmaniasis in the region. The rapid expansion of COVID-19 has further reduced resources for NTDs, exacerbated health inequality and reiterated the need to raise awareness of NTDs related to bites. Methods The nine countries that make up the Amazon basin have been considered (Bolivia, Brazil, Colombia, Ecuador, French Guiana, Guyana, Peru, Surinam and Venezuela) in the formation of a new network. Results The Amazonian Tropical Bites Research Initiative (ATBRI) has been created, with the aim of creating transdisciplinary solutions to the problem of animal bites leading to disease in Amazonian communities. The ATBRI seeks to unify the currently disjointed approach to the control of bite-related neglected zoonoses across Latin America. Conclusions The coordination of different sectors and inclusion of all stakeholders will advance this field and generate evidence for policy-making, promoting governance and linkage across a One Health arena.
Rabies is a fatal encephalitis caused by an important group of viruses within the Lyssavirus genus. The prototype virus, rabies virus, is still the most commonly reported lyssavirus and causes approximately 59,000 human fatalities annually. The human and animal burden of the other lyssavirus species is undefined. The original reports for the novel lyssavirus, Kotalahti bat lyssavirus (KBLV), were based on the detection of viral RNA alone. In this report we describe the successful generation of a live recombinant virus, cSN-KBLV; where the full-length genome clone of RABV vaccine strain, SAD-B19, was constructed with the glycoprotein of KBLV. Subsequent in vitro characterisation of cSN-KBLV is described here. In addition, the ability of a human rabies vaccine to confer protective immunity in vivo following challenge with this recombinant virus was assessed. Naïve or vaccinated mice were infected intracerebrally with a dose of 100 focus-forming units/30 µL of cSN-KBLV; all naïve mice and 8% (n = 1/12) of the vaccinated mice succumbed to the challenge, whilst 92% (n = 11/12) of the vaccinated mice survived to the end of the experiment. This report provides strong evidence for cross-neutralisation and cross-protection of cSN-KBLV using purified Vero cell rabies vaccine.
Many infectious diseases originating from or carried by wildlife impact wildlife conservation and biodiversity, livestock health, and/or human health. We provide an update on changes in the epidemiology of 25 selected infectious wildlife–related diseases in Europe (from 2010-2016) that had an impact, or may have a future impact, on the health of wildlife, livestock, and humans. These pathogens were selected based on their: (1) identification in recent Europe–wide projects as important surveillance targets, (2) inclusion in European Union (EU) legislation as pathogens requiring obligatory surveillance, (3) presence in recent literature on wildlife-related disease in Europe since 2010, (4) inclusion in key pathogen lists released by the Office International des Epizooties (OIE), (5) identification in conference presentations and informal discussions on a group email list by a European network of wildlife disease scientists from the European Wildlife Disease Association, or (6) identification as pathogens with changes in their epidemiology during 2010–2016. The wildlife pathogens or diseases included in this review are: avian influenza virus, seal influenza virus, lagoviruses, rabies virus, bat lyssaviruses, filoviruses, canine distemper virus, morbilliviruses in aquatic mammals, bluetongue virus, West Nile virus, hantaviruses, Schmallenberg virus, Crimean-Congo hemorrhagic fever virus, African swine fever virus, amphibian ranavirus, hepatitis E virus, bovine tuberculosis (Mycobacterium bovis), tularemia (Francisella tularensis), brucellosis (Brucella spp.), salmonellosis (Salmonella spp.), Coxiella burnetii , chytridiomycosis, Echinococcus multilocularis, Leishmania infantum, and chronic wasting disease. Further work is needed to identify all of the key drivers of disease change and emergence, as they appear to be influencing the incidence and spread of these pathogens in Europe. We present a summary of these recent changes over a specified time period to discuss possible commonalities and drivers of disease change and to identify directions for future work on wildlife related diseases in Europe. Many of the pathogens are entering Europe from other continents, while at the same time others are expanding their ranges inside and beyond Europe. Surveillance for these wildlife-related diseases at a continental scale is therefore important for planet-wide assessment, awareness of, and preparedness for, the potential risks they may pose to wildlife, domestic animal, and human health.
Background: The recommended screening of rabies in `suspect' animal cases involves testing fresh brain tissue. The preservation of fresh tissue however can be difficult under field conditions and formalin fixation provides a simple alternative that may allow a confirmatory diagnosis. The occurrence and location of histopathological changes and immunohistochemical (IHC) labelling for rabies in formalin fixed paraffin embedded (FFPE) canine brain is described in samples from 57 rabies suspect cases from Sri-Lanka. The presence of Negri bodies and immunohistochemical detection of rabies virus antigen were evaluated in the cortex, hippocampus, cerebellum and brainstem. The effect of autolysis and artefactual degeneration of the tissue was also assessed. Results: Rabies was confirmed in 53 of 57 (93%) cases by IHC. IHC labelling was statistically more abundant in the brainstem. Negri bodies were observed in 32 of 53 (60.4%) of the positive cases. Although tissue degradation had no effect on IHC diagnosis, it was associated with an inability to detect Negri bodies. In 13 cases, a confirmatory Polymerase chain reaction (PCR) testing for rabies virus RNA was undertaken by extracting RNA from fresh frozen tissue, and also attempted using FFPE samples. PCR detection using fresh frozen samples was in agreement with the IHC results. The PCR method from FFPE tissues was suitable for control material but unsuccessful in our field cases. Conclusions: Histopathological examination of the brain is essential to define the differential diagnoses of behaviour modifying conditions in rabies virus negative cases, but it is unreliable as the sole method for rabies diagnosis, particularly where artefactual change has occurred. Formalin fixation and paraffin embedding does not prevent detection of rabies virus via IHC labelling even where artefactual degeneration has occurred. This could represent a pragmatic secondary assay for rabies diagnosis in the field because formalin fixation can prevent sample degeneration. The brain stem was shown to be the site with most viral immunoreactivity; supporting recommended sampling protocols in favour of improved necropsy safety in the field. PCR testing of formalin fixed tissue may be successful in certain circumstances as an alternative test.
Rift Valley fever virus (RVFV) is a mosquito-borne arbovirus causing severe disease in humans and ruminants. Spread of RVFV out of Africa has raised concerns that it could emerge in Europe or the USA. Virus persistence is dependent on successful infection of, replication in, and transmission to susceptible vertebrate and invertebrate hosts, modulated by virus-host and vector-virus interactions. The principal accepted theory for the long term maintenance of RVFV involves vertical transmission (VT) of virus to mosquito progeny, with the virus surviving long inter-epizootic periods within the egg. This VT hypothesis however, is yet to be comprehensively proven. Here, evidence for and against the VT of RVFV is reviewed along with the identification of factors limiting its detection in natural and experimental data. The observation of VT for other arboviruses in the genera alphavirus, flavivirus and orthobunyavirus are discussed within the context of RVFV. The review concludes that VT of RVFV is likely but that current data are insufficient to irrefutably prove this hypothesis.
Lyssavirus infection has a near 100 % case fatality rate following the onset of clinical disease, and current rabies vaccines confer protection against all reported phylogroup I lyssaviruses. However, there is little or no protection against more divergent lyssaviruses and so investigation into epitopes within the glycoprotein (G) that dictate a neutralizing response against divergent lyssaviruses is warranted. Importantly, the facilities required to work with these pathogens, including wild-type and mutated forms of different lyssaviruses, are scarcely available and, as such, this type of study is inherently difficult to perform. The relevance of proposed immunogenic antigenic sites within the lyssavirus glycoprotein was assessed by swapping sites between phylogroup-I and -II glycoproteins. Demonstrable intra- but limited inter-phylogroup cross-neutralization was observed. Pseudotype viruses (PTVs) presenting a phylogroup-I glycoprotein containing phylogroup-II antigenic sites (I, II III or IV) were neutralized by antibodies raised against phylogroup-II PTV with the site II (IIb, aa 34-42 and IIa, aa 198-200)-swapped PTVs being efficiently neutralized, whilst site IV-swapped PTV was poorly neutralized. Specific antibodies raised against PTV-containing antigenic site swaps between phylogroup-I and -II glycoproteins neutralized phylogroup-I PTVs efficiently, indicating an immunodominance of antigenic site II. Live lyssaviruses containing antigenic site-swapped glycoproteins were generated and indicated that specific residues within the lyssavirus glycoprotein dictate functionality and enable differential neutralizing antibody responses to lyssaviruses.
Rift Valley fever virus (RVFV) causes a zoonotic mosquito-borne haemorrhagic disease that emerges to produce rapid large-scale outbreaks in livestock within sub-Saharan Africa. A range of mosquito species in Africa have been shown to transmit RVFV, and recent studies have assessed whether temperate mosquito species are also capable of transmission. In order to support vector competence studies, the ability to visualize virus localization in mosquito cells and tissue would enhance the understanding of the infection process within the mosquito body. Here, the application of in situ hybridization utilizing RNAscope® to detect RVFV infection within the mosquito species, Culex pipiens, derived from the United Kingdom was demonstrated. Extensive RVFV replication was detected in many tissues of the mosquito with the notable exception of the interior of ovarian follicles.
Pathogens causing acute disease and death or lasting immunity require specific spatial or temporal processes to persist in populations. Host traits, such as maternally-derived antibody (MDA) and seasonal birthing affect infection maintenance within populations. Our study objective is to understand how viral and host traits lead to population level infection persistence when the infection can be fatal. We collected data on African fruit bats and a rabies-related virus, Lagos bat virus (LBV), including through captive studies. We incorporate these data into a mechanistic model of LBV transmission to determine how host traits, including MDA and seasonal birthing, and viral traits, such as incubation periods, interact to allow fatal viruses to persist within bat populations. Captive bat studies supported MDA presence estimated from field data. Captive bat infection-derived antibody decayed more slowly than MDA, and while faster than estimates from the field, supports field data that suggest antibody persistence may be lifelong. Unobserved parameters were estimated by particle filtering and suggest only a small proportion of bats die of disease. Pathogen persistence in the population is sensitive to this proportion, along with MDA duration and incubation period. Our analyses suggest MDA produced bats and prolonged virus incubation periods allow viral maintenance in adverse conditions, such as a lethal pathogen or strongly seasonal resource availability for the pathogen in the form of seasonally pulsed birthing.
Background: Rift Valley fever phlebovirus (RVFV) is a mosquito-borne arbovirus causing severe disease in humans and livestock. It is endemic in Africa and spread to the Arabian Peninsula in 2000 raising concerns it could emerge in Europe. The ability of temperate mosquitoes from the United Kingdom (UK) to support replication and transmission of RVFV is unknown. Methods: In this study, two colonised lines of Culex pipiens, wild-caught Aedes detritus and Ae. rusticus from the UK were infected with pathogenic strains of RVFV to assess their vector competence. Mosquitoes were offered artificial blood-meals containing 10(6) or 10(7) plaque forming units (PFU)/ml RVFV, simulating natural peak viraemia in young ruminants, and maintained at 20 degrees C or 25 degrees C for up to 21 days. Bodies, legs and saliva were collected and tested for the presence of viral RNA and infectious virus to determine the infection, dissemination and transmission potential. Results: Across temperatures, doses and strains the average infection, dissemination and transmission rates were: 35, 13 and 5% (n = 91) for Cx. pipiens (Caldbeck); 23, 14 and 5% (n = 138) for Cx. pipiens (Brookwood); 36, 28 and 7% (n = 118) for Ae. detritus. However, despite 35% (n = 20) being susceptible to infection, Ae rusticus did not transmit RVFV. Survival of Aedes species was negatively affected by maintenance at 25 degrees C compared to the more representative peak average British summer temperature of 20 degrees C. Increased mortality was also observed with some species infected with 10(7) PFU/ml compared to 10(6 )PFU/ml. Conclusions: It can be concluded that temperate mosquito species present in the UK demonstrate a transmission potential for RVFV in the laboratory but, even at high temperatures, this occurred at low efficiency.
Here we report studies of the antigenic relationship of West Nile virus (WNV) and Usutu virus (USUV), two zoonotic flaviviruses from Italy, together with a Japanese encephalitis virus (JEV) strain and compared them with their genetic relationship using the immunodominant viral E protein. Thirty-nine isolates and reference strains were inactivated and used to immunize rabbits to produce hyper immune sera. Serum samples were tested by neutralization against all isolates and results visualized by generating antigenic map. Strains of WNV, USUV, and JEV grouped in separate clusters on the antigenic map. JEV was closer antigenically to USUV (mean of 3.5 Antigenic Unit, AU, equivalent to a 2-fold change in antibody titer) than to WNV strains (mean of 6 AU). A linear regression model predicted, on average, one unit of antigenic change, equivalent to a 2-fold change in antibody titer, for every 22 amino acid substitutions in the E protein ectodomain. Overall, antigenic map was demonstrated to be robust and consistent with phylogeny of the E protein. Indeed, the map provided a reliable means of visualizing and quantifying the relationship between these flaviviruses. Further antigenic analyses employing representative strains of extant serocomplexes are currently underway. This will provide a more in deep knowledge of antigenic relationships between flaviviruses.
In Nepal most dogs are free to roam and may transmit diseases to humans and animals. These dogs often suffer from malnutrition and lack basic health care. Minimal information is available about their demographics and about public attitudes concerning dogs and diseases. We carried out a study in Chitwan District (central Nepal), to collect baseline data on free-roaming owned dog demographics, assess knowledge, attitudes and practices of dog owners concerning dogs and rabies, evaluate rabies vaccination coverage and anthelmintic treatment of dogs, measure dogs’ response to rabies vaccination and assess dog health through body condition scores and parasites. We conducted household interviews with owners of free-roaming female dogs (n=60) and administered dogs with rabies vaccination and anthelmintics. Dog owners regularly fed free-roaming dogs but provided minimal health care; 42% of respondents did not claim ownership of the dog for which they provided care. We collected skin, faecal and blood samples for parasite identification and for measuring rabies virus-specific antibodies. Ninety-two percent of dog owners were aware of the routes of rabies virus transmission but only 35% described the correct post-exposure prophylaxis (PEP) following a dog bite. Twenty-seven percent of the dogs had measurable rabies virus-specific antibody titres and 14% had received anthelmintics in the previous year. Following rabies vaccination, 97% of dogs maintained an adequate antibody titre for > 6 months. Most dogs appeared healthy, although haemoprotozoans, endoparasites and ectoparasites were identified in 12%, 73% and 40% of the dogs, respectively. Poor skin condition and parasite load were associated. Seventy-four percent of the females had litters in one year (mean litter size= 4.5). Births occurred between September and February; we estimated 60% mortality in puppies. We concluded that vaccination coverage, PEP awareness and anthelmintic treatment should be emphasized in educational programmes focussed on animal welfare, veterinary and public health.
Many high-consequence human and animal pathogens persist in wildlife reservoirs. An understanding of the dynamics of these pathogens in their reservoir hosts is crucial to inform the risk of spill-over events, yet our understanding of these dynamics is frequently insufficient. Viral persistence in a wild bat population was investigated by combining empirical data and in-silico analyses to test hypotheses on mechanisms for viral persistence. A fatal zoonotic virus, European Bat lyssavirus type 2 (EBLV-2), in Daubenton's bats (Myotis daubentonii) was used as a model system. A total of 1839 M. daubentonii were sampled for evidence of virus exposure and excretion during a prospective nine year serial cross-sectional survey. Multivariable statistical models demonstrated age-related differences in seroprevalence, with significant variation in seropositivity over time and among roosts. An Approximate Bayesian Computation approach was used to model the infection dynamics incorporating the known host ecology. The results demonstrate that EBLV-2 is endemic in the study population, and suggest that mixing between roosts during seasonal swarming events is necessary to maintain EBLV-2 in the population. These findings contribute to understanding how bat viruses can persist despite low prevalence of infection, and why infection is constrained to certain bat species in multispecies roosts and ecosystems.
In South Africa, canid rabies virus (RABV) infection is maintained in domestic and wildlife species. The identification of rabies in African civets raised the question of whether this wildlife carnivore is a potential reservoir host of RABVs of direct and ancestral dog origin (dog-maintained and dog-derived origins) with an independent cycle of transmission. Genetic analyses of African civet nucleoprotein sequences for 23 African civet RABVs and historically published sequences demonstrated that RABVs from African civets have two origins related to dog and mongoose rabies enzootics. The data support observations of the interaction of civets with domestic dogs and wildlife mongooses, mostly in Northern South Africa and North-East Zimbabwe. Within each host species clade, African civet RABVs group exclusively together, implying intra-species virus transfer occurs readily. The canid RABV clade appears to support virus transfer more readily between hosts than mongoose RABVs. Furthermore, these data probably indicate short transmission chains with conspecifics that may be related to transient rabies maintenance in African civets. Hence, it is important to continue monitoring the emergence of lyssaviruses in this host. Observations from this study are supported by ongoing and independent similar cases, in which bat-eared foxes and black-backed jackal species maintain independent rabies cycles of what were once dog-maintained RABVs.
A brain homogenate derived from a rabid dog in the district of Tojiko-bod, Republic of Tajikistan, was applied to a Flinders Technology Associates (FTA) card. A full-genome sequence of rabies virus (RABV) was generated from the FTA card directly without extraction, demonstrating the utility of these cards for readily obtaining genetic data.
Many of the pathogens perceived to pose the greatest risk to humans are viral zoonoses, responsible for a range of emerging and endemic infectious diseases. Phylogeography is a useful tool to understand the processes that give rise to spatial patterns and drive dynamics in virus populations. Increasingly, whole-genome information is being used to uncover these patterns, but the limits of phylogenetic resolution that can be achieved with this are unclear. Here, whole-genome variation was used to uncover fine-scale population structure in endemic canine rabies virus circulating in Tanzania. This is the first whole-genome population study of rabies virus and the first comprehensive phylogenetic analysis of rabies virus in East Africa, providing important insights into rabies transmission in an endemic system. In addition, sub-continental scale patterns of population structure were identified using partial gene data and used to determine population structure at larger spatial scales in Africa. While rabies virus has a defined spatial structure at large scales, increasingly frequent levels of admixture were observed at regional and local levels. Discrete phylogeographic analysis revealed long-distance dispersal within Tanzania, which could be attributed to human-mediated movement, and we found evidence of multiple persistent, co-circulating lineages at a very local scale in a single district, despite on-going mass dog vaccination campaigns. This may reflect the wider endemic circulation of these lineages over several decades alongside increased admixture due to human-mediated introductions. These data indicate that successful rabies control in Tanzania could be established at a national level, since most dispersal appears to be restricted within the confines of country borders but some coordination with neighbouring countries may be required to limit transboundary movements. Evidence of complex patterns of rabies circulation within Tanzania necessitates the use of whole-genome sequencing to delineate finer scale population structure that can that can guide interventions, such as the spatial scale and design of dog vaccination campaigns and dog movement controls to achieve and maintain freedom from disease.
SARS-CoV-2 variants may threaten the effectiveness of vaccines and antivirals to mitigate serious COVID-19 disease. This is of most concern in clinically vulnerable groups such as older adults. We analysed 72 sera samples from 37 individuals, aged 70–89 years, vaccinated with two doses of BNT162b2 (Pfizer–BioNTech) 3 weeks apart, for neutralizing antibody responses to wildtype SARS-CoV-2. Between 3 and 20 weeks after the second vaccine dose, neutralizing antibody titres fell 4.9-fold to a median titre of 21.3 (neutralization dose 80%), with 21.6% of individuals having no detectable neutralizing antibodies at the later time point. Next, we examined neutralization of 21 distinct SARS-CoV-2 variant spike proteins with these sera, and confirmed substantial antigenic escape, especially for the Omicron (B.1.1.529, BA.1/BA.2), Beta (B.1.351), Delta (B.1.617.2), Theta (P.3), C.1.2 and B.1.638 spike variants. By combining pseudotype neutralization with specific receptor-binding domain (RBD) enzyme-linked immunosorbent assays, we showed that changes to position 484 in the spike RBD were mainly responsible for SARS-CoV-2 neutralizing antibody escape. Nineteen sera from the same individuals boosted with a third dose of BNT162b2 contained higher neutralizing antibody titres, providing cross-protection against Omicron BA.1 and BA.2. Despite SARS-CoV-2 immunity waning over time in older adults, booster vaccines can elicit broad neutralizing antibodies against a large number of SARS-CoV-2 variants in this clinically vulnerable cohort. Analysis of the neutralizing antibody activity from sera of vaccinated individuals aged between 70 and 89 reveals a reduction of antibody titres against SARS-CoV-2 wildtype and antigenic escape of various variants of concern that links to specific mutations within the RBD. A booster vaccination helps increasing neutralizing antibody titres against the Omicron BA.1 and BA.2 variants in older adults.
The mountain chicken Leptodactylus fallax, the largest amphibian in the Caribbean, restricted to Dominica and Montserrat, is Critically Endangered. We investigated the presence of disease in the mountain chicken and in the sympatric cane toad Bufo marinus and Johnstone's whistling frog Eleutherodactylus johnstonei in the Centre Hills on Montserrat. Skin swabs of mountain chickens and B. marinus, and toe clips from E. johnstonei, were tested for Batrachochytrium dendrobatidis, the causative organism of cutaneous chytridiomycosis. Additionally, mountain chicken blood serum samples (n = 57) were examined for exposure to ranavirus. No chytrid or exposure to ranavirus was detected. We draw tentative conclusions about disease threats to the Montserrat mountain chicken population, and present preliminary recommendations for safeguarding this species.
Effective methods to increase awareness of preventable infectious diseases are key components of successful control programs. Rabies is an example of a disease with significant impact, where public awareness is variable. A recent awareness campaign in a rabies endemic region of Azerbaijan provided a unique opportunity to assess the efficacy of such campaigns. A cluster-cross sectional survey concerning rabies was undertaken following the awareness campaign in 600 households in 38 randomly selected towns, in districts covered by the campaign and matched control regions. This survey demonstrated that the relatively simple awareness campaign was effective at improving knowledge of rabies symptoms and vaccination schedules. Crucially, those in the awareness campaign group were also 1·4 times more likely to report that they had vaccinated their pets, an essential component of human rabies prevention. In addition, low knowledge of appropriate post-exposure treatment and animal sources of rabies provide information useful for future public awareness campaigns in the region and other similar areas.
Using next-generation sequencing technologies, the first complete genome sequence of Rift Valley fever virus strain Lunyo is reported here. Originally reported as an attenuated antigenic variant strain from Uganda, genomic sequence analysis shows that Lunyo clusters together with other Ugandan isolates.
Disease surveillance in wildlife populations presents a logistical challenge, yet is critical in gaining a deeper understanding of the presence and impact of wildlife pathogens. Erinaceus coronavirus (EriCoV), a clade C Betacoronavirus, was first described in Western European hedgehogs (Erinaceus europaeus) in Germany. Here, our objective was to determine whether EriCoV is present, and if it is associated with disease, in Great Britain (GB). An EriCoV-specific BRYT-Green® real-time reverse transcription PCR assay was used to test 351 samples of faeces or distal large intestinal tract contents collected from casualty or dead hedgehogs from a wide area across GB. Viral RNA was detected in 10.8% (38) samples; however, the virus was not detected in any of the 61 samples tested from Scotland. The full genome sequence of the British EriCoV strain was determined using next generation sequencing; it shared 94% identity with a German EriCoV sequence. Multivariate statistical models using hedgehog case history data, faecal specimen descriptions and post-mortem examination findings found no significant associations indicative of disease associated with EriCoV in hedgehogs. These findings indicate that the Western European hedgehog is a reservoir host of EriCoV in the absence of apparent disease.
Host shift events play an important role in epizootics as adaptation to new hosts can profoundly affect the spread of the disease and the measures needed to control it. During the late 1990s, an epizootic in Turkey resulted in a sustained maintenance of rabies virus (RABV) within the fox population. Utilisation of Bayesian inferences to investigate whole genome sequences from a cohort of fox and dog brain tissues from Turkey demonstrated that the epizootic occurred in 1997 (+/- 1 year). Furthermore, these data indicate that the epizootic was most likely due to a host shift from locally infected domestic dogs, rather than an incursion of a novel fox or dog RABV. No evidence was detected for virus adaptation to foxes at consensus sequence level; therefore, the deep sequence data was analysed to investigate the influence of sub-consensus populations on host shift events. Viral heterogeneity was measured in all RABV samples; viruses in the early phase after the host shift had increased heterogeneity, in relation to those in the later stage, possibly indicating a role in establishing transmission within a new host. The dynamics of majority and minority variants are consistent with genetic drift, rather than positive selection. The transient expansion of sub-consensus viral populations in the new host species likely represents the virus adapting to a new environment, perhaps due to increased replication within the CNS resulting in a larger population of viruses, or reflecting the lack of host constraints present in the new host reservoir.
Bluetongue (BT) is a severe and economically important disease of ruminants that is widely distributed around the world, caused by the bluetongue virus (BTV). More than 28 different BTV serotypes have been identified in serum neutralisation tests (SNT), which, along with geographic variants (topotypes) within each serotype, reflect differences in BTV outer-capsid protein VP2. VP2 is the primary target for neutralising antibodies, although the basis for cross-reactions and serological variations between and within BTV serotypes is poorly understood. Recombinant BTV VP2 proteins (rVP2) were expressed in Nicotiana benthamiana, based on sequence data for isolates of thirteen BTV serotypes (primarily from Europe), including three ‘novel’ serotypes (BTV-25, -26 and -27) and alternative topotypes of four serotypes. Cross-reactions within and between these viruses were explored using rabbit anti-rVP2 sera and post BTV-infection sheep reference-antisera, in I-ELISA (with rVP2 target antigens) and SNT (with reference strains of BTV-1 to -24, -26 and -27). Strong reactions were generally detected with homologous rVP2 proteins or virus strains/serotypes. The sheep antisera were largely serotype-specific in SNT, but more cross-reactive by ELISA. Rabbit antisera were more cross-reactive in SNT, and showed widespread, high titre cross-reactions against homologous and heterologous rVP2 proteins in ELISA. Results were analysed and visualised by antigenic cartography, showing closer relationships in some, but not all cases, between VP2 topotypes within the same serotype, and between serotypes belonging to the same ‘VP2 nucleotype’.